Improve your Predictive Model’s Score using a Stacking Regressor
This article was published as a part of the Data Science Blogathon.
Introduction
The general principle of ensembling is to combine the predictions of various models built with a given learning algorithm in order to improve robustness over a single model.
“The whole is greater than the sum of its parts.” – Aristotle
In other words, when individual parts are connected together to form one entity, they are worth more than if the parts were separate.
The above quote is more apt for stacking where we combine different models to get better performance. In this article, we will discuss stacking and also how to create your own stacking regressor.
What is meant by stacking?
Ensemble learning is a technique widely used by machine learning practitioners, that combines the skills of different models to make predictions from the given data. We are using this to combine the best of multiple algorithms that can give more stable predictions with very little variance than what we get with a single regressor.
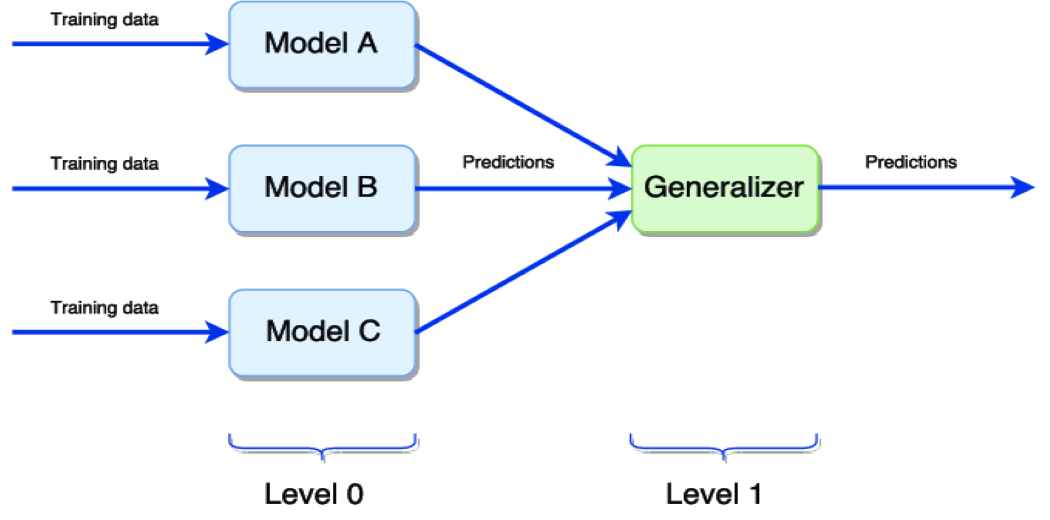
The above diagram represents the simple stacking of the models,
Level 0 – Training different models on the same dataset then making predictions.
Level 1 – Generalize the predictions made by different models to get the final output.
The most common way of generalizer is by taking the average of all the level 0 model predictions to get the final output.
Creating your own stacking regressor
We will take the famous House price prediction dataset from Kaggle. Where the objective is to predict the house price based on several features that are present in the dataset.
Base models
We will train different base-models using k-fold cross-validation and see the performance(RMSE) of all the models in the given dataset.
The below function rmse_cv is used to train all the individual models in the 5 folds of the data created and it returns the RMSE score for the model based on the out of fold predictions compared with the actual predictions.
Note: All the Data preprocessing techniques have been done before training the base models.
Lasso
lasso = Lasso(alpha =0.0005) rmse_cv(lasso)
Lasso score: 0.1115
ElasticNet
ENet = ElasticNet(alpha=0.0005, l1_ratio=.9) rmse_cv(ENet)
ElasticNet Score: 0.1116
Kernel Ridge Regression
KRR = KernelRidge(alpha=0.6, kernel='polynomial', degree=2, coef0=2.5) rmse_cv(KRR)
Kernel Ridge Score: 0.1153
Gradient Boosting
GBoost = GradientBoostingRegressor(n_estimators=3000, learning_rate=0.05,max_depth=4,max_features='sqrt',min_samples_leaf=15, min_samples_split=10,loss='huber') rmse_cv(GBoost)
Gradient Boosting Score: 0.1177
XGB Regressor
model_xgb = xgb.XGBRegressor(colsample_bytree=0.4603, gamma=0.0468,
learning_rate=0.05, max_depth=3,
min_child_weight=1.7817, n_estimators=2200,
reg_alpha=0.4640, reg_lambda=0.8571,
subsample=0.5213, silent=1,
random_state =7, nthread = -1)
rmse_cv(model_xgb)
XGBoost Score: 0.1161
LightGBM
model_lgb = lgb.LGBMRegressor(objective='regression',num_leaves=5,
learning_rate=0.05, n_estimators=720,
max_bin = 55, bagging_fraction = 0.8,
bagging_freq = 5, feature_fraction = 0.2319,
feature_fraction_seed=9, bagging_seed=9,
min_data_in_leaf =6, min_sum_hessian_in_leaf = 11)
rmse_cv(model_lgb)
LGBM Score: 0.1157
Type 1: Simplest Stacking Regressor approach: Averaging Base models
We begin with this simple approach of averaging base models. Build a new class to extend scikit-learn with our model and also to leverage encapsulation and code reuse.
Averaged base models class
from sklearn.base import BaseEstimator, TransformerMixin, RegressorMixin, clone
class AveragingModels(BaseEstimator, RegressorMixin, TransformerMixin):
def __init__(self, models):
self.models = models
# we define clones of the original models to fit the data in
def fit(self, X, y):
self.models_ = [clone(x) for x in self.models]
# Train cloned base models
for model in self.models_:
model.fit(X, y)
return self
#Now we do the predictions for cloned models and average them
def predict(self, X):
predictions = np.column_stack([
model.predict(X) for model in self.models_
])
return np.mean(predictions, axis=1)
fit() – This method clones all the base models which were passed as a parameter, and then train the cloned models in the whole dataset.
predict() – All the cloned models will predict and the predictions will be column stacked using “np.column_stack()” then the average of that prediction will be calculated and returned.
averaged_models = AveragingModels(models = (ENet, GBoost, KRR, lasso))
score = rmsle_cv(averaged_models)
print(" Averaged base models score: {:.4f}n".format(score.mean()))
Averaged base models score: 0.1091
Wow…..! Even simply taking the average of all the base models predictions gave us a good RMSE score.
Type 2: Adding a Meta-model
In this approach, we will train all the base models and use the predictions (out-of-fold predictions) of the base models as a training feature to the meta-model.
The meta-model is used to find the pattern between the base model predictions as features and actual predictions as the target variables.
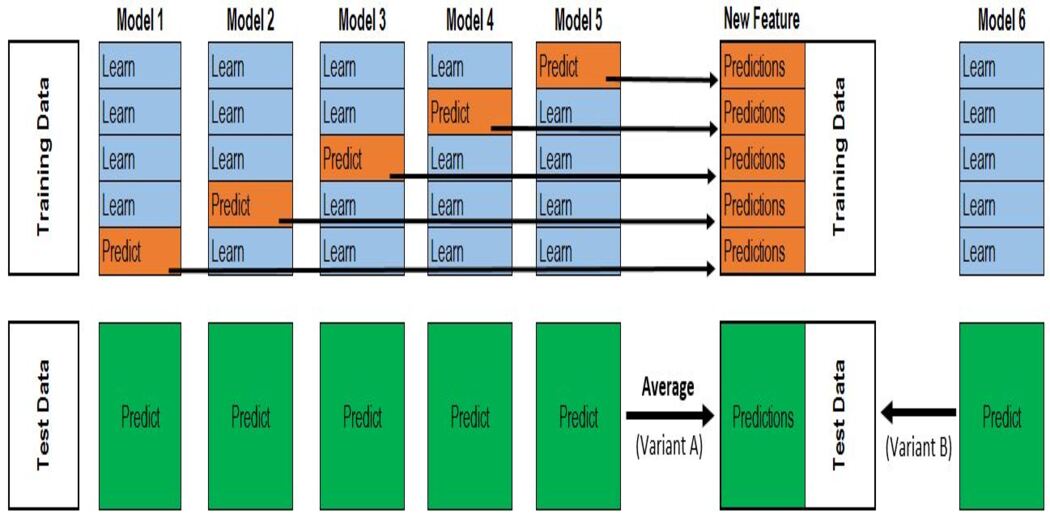
Steps:
1. Split the data into 2 sets training and holdout set.
2. Train all the base models in the training data.
3. Test base models on the holdout dataset and store the predictions(out-of-fold predictions).
4. Use the out-of-fold predictions made by the base models as input features, and the correct output as the target variable to train the meta-model.
The first three steps will be done iteratively for the k-folds based on the value of k. If k=5 then we will train the model on the 4 folds and predict on the holdout set (5th fold). Repeating this step for k-times (here,k=5) gives the out-of-fold predictions for the whole dataset. This will be done for all the base models.
Then meta-model will be trained using the out-of-predictions by all the models as X and the original target variable as y. Prediction of this meta-model will be considered as the final prediction.
class StackingAveragedModels(BaseEstimator, RegressorMixin, TransformerMixin):
def __init__(self, base_models, meta_model, n_folds=5):
self.base_models = base_models
self.meta_model = meta_model
self.n_folds = n_folds
# We again fit the data on clones of the original models
def fit(self, X, y):
self.base_models_ = [list() for x in self.base_models]
self.meta_model_ = clone(self.meta_model)
kfold = KFold(n_splits=self.n_folds, shuffle=True, random_state=156)
# Train cloned base models then create out-of-fold predictions
# that are needed to train the cloned meta-model
out_of_fold_predictions = np.zeros((X.shape[0], len(self.base_models)))
for i, model in enumerate(self.base_models):
for train_index, holdout_index in kfold.split(X, y):
instance = clone(model)
self.base_models_[i].append(instance)
instance.fit(X[train_index], y[train_index])
y_pred = instance.predict(X[holdout_index])
out_of_fold_predictions[holdout_index, i] = y_pred
# Now train the cloned meta-model using the out-of-fold predictions as new feature
self.meta_model_.fit(out_of_fold_predictions, y)
return self
#Do the predictions of all base models on the test data and use the averaged predictions as
#meta-features for the final prediction which is done by the meta-model
def predict(self, X):
meta_features = np.column_stack([
np.column_stack([model.predict(X) for model in base_models]).mean(axis=1)
for base_models in self.base_models_ ])
return self.meta_model_.predict(meta_features)
fit() – Cloning the base model and meta-model, Training the base model in 5fold cross-validation, and storing the out of fold predictions. Then training the meta-model using the out-of-fold-predictions.
predict() – base model predictions for X will be column stacked and then used as an input for meta-model to predict.
stacked_averaged_models = StackingAveragedModels(base_models = (ENet, GBoost, KRR),
meta_model = lasso)
score = rmsle_cv(stacked_averaged_models)
print("Stacking Averaged models score: {:.4f}".format(score.mean()))
Stacking Averaged models score: 0.1085
Whoa…!!! We again got a better score by using the meta learner.
Conclusion
The above stacking method can also be used for the classification task with some minor changes. Instead of averaging the base model predictions, we can do voting of all the model predictions and the highest voted class can be taken as an output.
This notebook shows an example of using the above method from data preprocessing to model building.
Did you find this article useful? Share your views and opinions in the comment section









Thank you for your interesting article. It contains not only useful Stacking Approach but useful review of different regressions used in modelling.