A Brief Introduction to Linear Discriminant Analysis
This article was published as a part of the Data Science Blogathon
Introduction to LDA:
Linear Discriminant Analysis as its name suggests is a linear model for classification and dimensionality reduction. Most commonly used for feature extraction in pattern classification problems. This has been here for quite a long time. First, in 1936 Fisher formulated linear discriminant for two classes, and later on, in 1948 C.R Rao generalized it for multiple classes. LDA projects data from a D dimensional feature space down to a D’ (D>D’) dimensional space in a way to maximize the variability between the classes and reducing the variability within the classes.
Table of contents
What is Linear Discriminant Analysis?
Linear Discriminant Analysis (LDA) is a statistical technique for categorizing data into groups. It identifies patterns in features to distinguish between different classes. For instance, it may analyze characteristics like size and color to classify fruits as apples or oranges. LDA aims to find a straight line or plane that best separates these groups while minimizing overlap within each class. By maximizing the separation between classes, it enables accurate classification of new data points. In simpler terms, LDA helps make sense of data by finding the most effective way to separate different categories, aiding tasks like pattern recognition and classification.
Why Linear Discriminant Analysis (LDA)?:
- Logistic Regression is one of the most popular linear classification models that perform well for binary classification but falls short in the case of multiple classification problems with well-separated classes. While LDA handles these quite efficiently.
- Linear Discriminant Analysis (LDA) also reduces the number of features in data preprocessing, akin to Principal Component Analysis (PCA), thereby significantly reducing computing costs.
- LDA is also used in face detection algorithms. In Fisherfaces LDA is used to extract useful data from different faces. Coupled with eigenfaces it produces effective results.
Shortcomings:
- Linear decision boundaries may not effectively separate non-linearly separable classes. More flexible boundaries are desired.
- In cases where the number of observations exceeds the number of features, LDA might not perform as desired. This is called Small Sample Size (SSS) problem. Regularization is required.
We will discuss this later.
Assumptions:
Linear Discriminant Analysis (LDA) makes some assumptions about the data:
- It assumes that the data follows a normal or Gaussian distribution, meaning each feature forms a bell-shaped curve when plotted.
- Each of the classes has identical covariance matrices.
However, it is worth mentioning that LDA performs quite well even if the assumptions are violated.
Fisher’s Linear Discriminant:
Linear Discriminant Analysis (LDA) is a generalized form of FLD. Fisher in his paper used a discriminant function to classify between two plant species Iris Setosa and Iris Versicolor.
The basic idea of FLD is to project data points onto a line to maximize the between-class scatter and minimize the within-class scatter.
This might sound a bit cryptic but it is quite straightforward. So, before delving deep into the derivation part we need to get familiarized with certain terms and expressions.
- Let’s suppose we have d-dimensional data points x1….xn with 2 classes Ci=1,2 each having N1 & N2 samples.
- Let W be a unit vector onto which the data points are to be projected (took unit vector as we are only concerned with the direction).
- Number of samples : N = N1 + N2
- If x(n) are the samples on the feature space then WTx(n) denotes the data points after projection.
- Means of classes before projection: mi
- Means of classes after projection: Mi = WTmi
.png)
Scatter matrix: Used to make estimates of the covariance matrix. IT is a m X m positive semi-definite matrix.
Given by: sample variance * no. of samples.
Note: Scatter and variance measure the same thing but on different scales. So, we might use both words interchangeably. So, do not get confused.
Two Types of Scatter Matrices
Here we will be dealing with two types of scatter matrices
- Between class scatter = Sb = measures the distance between class means
- Within class scatter = Sw = measures the spread around means of each class
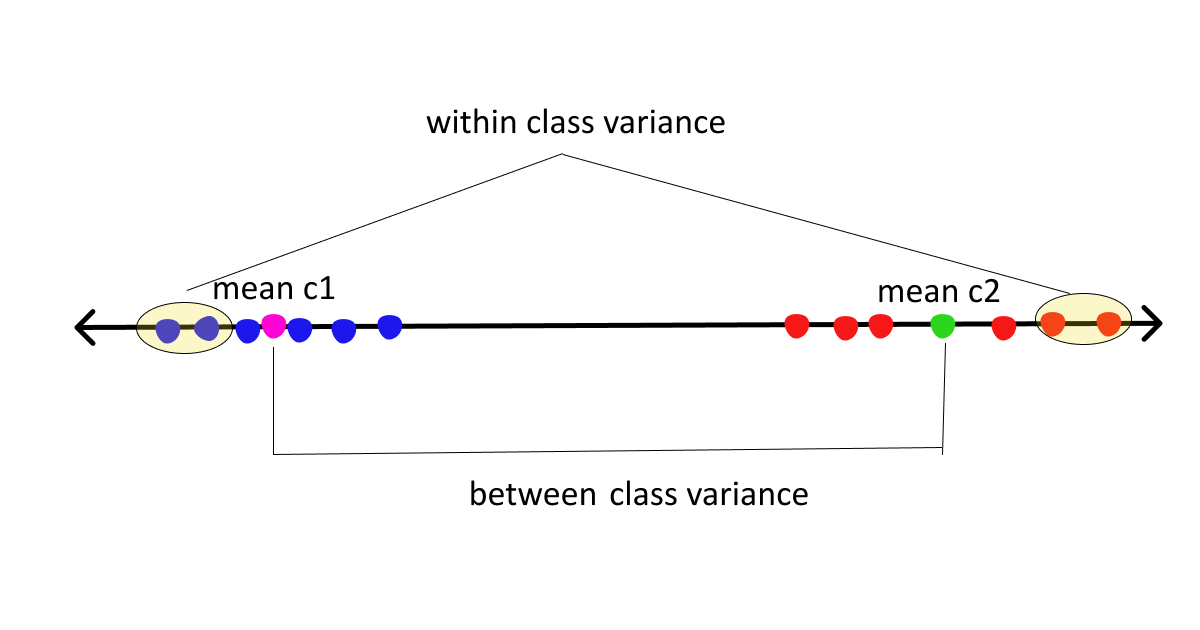
Now, assuming we are clear with the basics let’s move on to the derivation part.
As per Fisher’s LDA :
arg max J(W) = (M1 – M2)2 / S12 + S22 ……….. (1)
The numerator here is between class scatter while the denominator is within-class scatter. So to maximize the function we need to maximize the numerator and minimize the denominator, simple math. To maximize the above function we need to first express the above equation in terms of W.
Numerator:
.jpg)
Denominator:
For denominator we have S12 + S22 .

Now, we have both the numerator and denominator expressed in terms of W
J(W) = WTSbW / WTSwW
Upon differentiating the above function w.r.t W and equating with 0, we get a generalized eigenvalue-eigenvector problem
SbW = vSwW
Sw being a full-rank matrix , inverse is feasible
=> Sw-1SbW = vW
Where v = eigen value
W = eigen vector
Linear Discriminant Analysis (LDA) for multiple classes:
Linear Discriminant Analysis (LDA) can be generalized for multiple classes. Here are the generalized forms of between-class and within-class matrices.
.png)
Note: Sb is the sum of C different rank 1 matrices. So, the rank of Sb <=C-1. That means we can only have C-1 eigenvectors. Thus, we can project data points to a subspace of dimensions at most C-1.
Above equation (4) gives us scatter for each of our classes and equation (5) adds all of them to give within-class scatter. Similarly, equation (6) gives us between-class scatter. Finally, eigendecomposition of Sw-1Sb gives us the desired eigenvectors from the corresponding eigenvalues. Total eigenvalues can be at most C-1.
LDA for classification:
Until now, we only reduced the dimension of the data points, but this is strictly not yet discriminant. But the projected data can subsequently be used to construct a discriminant by using Bayes’ theorem as follows.
Assume X = (x1….xp) is drawn from a multivariate Gaussian distribution. K be the no. of classes and Y is the response variable. pik is the prior probability: the probability that a given observation is associated with Kth class. Pr(X = x | Y = k) is the posterior probability.
Let fk(X) = Pr(X = x | Y = k) is our probability density function of X for an observation x that belongs to Kth class. fk(X) is large if there is a high probability of an observation in Kth class has X=x.
As per Bayes’ theorem,
.png)
Now, to calculate the posterior probability we will need to find the prior pik and density function fk(X).
pik can be calculated easily. If we have a random sample of Ys from the population: we simply compute the fraction of the training observations that belong to Kth class. But the calculation of fk(X) can be a little tricky.
We assume that the probability density function of x is multivariate Gaussian with class means mk and a common covariance matrix sigma.
As a formula, multi-variate Gaussian density is given by:
.png)
|sigma| = determinant of covariance matrix ( same for all classes)
mk = class means
Now, by plugging the density function in the equation (8), taking the logarithm and doing some algebra, we will find the Linear score function
.png)
We will classify a sample unit to the class that has the highest Linear Score function for it.
Note that in the above equation (9) Linear discriminant function depends on x linearly, hence the name Linear Discriminant Analysis.
Addressing LDA shortcomings
Linearity problem: Linear Discriminant Analysis (LDA) is used to find a linear transformation that classifies different classes. But if the classes are non-linearly separable, It can not find a lower-dimensional space to project. This problem arises when classes have the same means i.e, the discriminatory information does not exist in mean but in the scatter of data. That will effectively make Sb=0. To address this issue we can use Kernel functions. As used in SVM, SVR etc. The idea is to map the input data to a new high dimensional feature space by a non-linear mapping where inner products in the feature space can be computed by kernel functions.
Small Sample problem: This problem arises when the dimension of samples is higher than the number of samples (D>N). This is the most common problem with LDA. The covariance matrix becomes singular, hence no inverse. So, to address this problem regularization was introduced. Instead of using sigma or the covariance matrix directly, we use
.png)
Here, alpha is a value between 0 and 1.and is a tuning parameter. i is the identity matrix. The diagonal elements of the covariance matrix are biased by adding this small element. However, the regularization parameter needs to be tuned to perform better.
In discriminant analysis machine learning, Scikit Learn’s Linear Discriminant Analysis incorporates a shrinkage parameter to tackle undersampling issues. This parameter enhances the classifier’s generalization performance. When set to ‘auto’, it automatically determines the best shrinkage parameter. Note that it operates only when the solver parameter is ‘lsqr’ or ‘eigen’. Users can manually adjust this parameter between 0 and 1. Additionally, various other methods address undersampling, such as combining PCA and LDA. PCA reduces dimensions first, followed by regular LDA machine learning procedures.
Python Implementation:
Fortunately, we don’t have to code all these things from scratch, Python has all the necessary requirements for LDA implementations. For the following article, we will use the famous wine dataset.
Python Code:
Fitting LDA to wine dataset:
lda = LinearDiscriminantAnalysis()
lda_t = lda.fit_transform(X,y)
Variance explained by each component:
lda.explained_variance_ratio_
.png)
Plotting LDA components:
plt.xlabel('LD1')
plt.ylabel('LD2')
plt.scatter(lda_t[:,0],lda_t[:,1],c=y,cmap='rainbow',edgecolors='r')
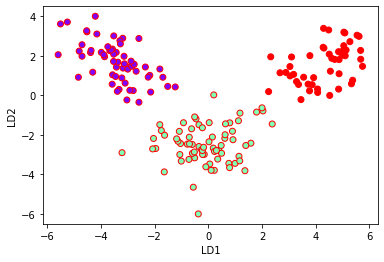
LDA for classification:
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size=0.3)
lda.fit(X_train,y_train)Accuracy Score:
y_pred = lda.predict(X_test)
print(accuracy_score(y_test,y_pred))
.png)
Confusion Matrix:
confusion_matrix(y_test,y_pred)
.png)
Plotting Decision boundary for our dataset:
min1,max1 = lda_t[:,0].min()-1, lda_t[:,0].max()+1
min2,max2 = lda_t[:,1].min()-1,lda_t[:,1].max()+1
x1grid = np.arange(min1,max1,0.1)
x2grid = np.arange(min2,max2,0.1)
xx,yy = np.meshgrid(x1grid,x2grid)
r1,r2 = xx.flatten(),yy.flatten()
r1,r2 = r1.reshape((len(r1),1)), r2.reshape((len(r2),1))
grid = np.hstack((r1,r2))
model = LinearDiscriminantAnalysis()
model.fit(lda_t,y)
yhat = model.predict(grid)
zz = yhat.reshape(xx.shape)
plt.contourf(xx,yy,zz,cmap='Accent')
for class_value in range(3):
row_ix = np.where( y== class_value)
plt.scatter(lda_t[row_ix,0],lda_t[row_ix,1])
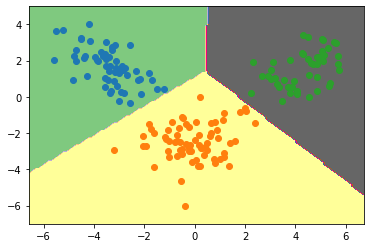
Conclusion
Linear Discriminant Analysis (LDA) stands out as a valuable tool for simplifying data and making classifications. We’ve covered why LDA is important, its limitations, and key components such as Fisher’s Linear Discriminant. Understanding the numerator and denominator in LDA, its application for multiple classes, and its implementation in Python provides practical insights. Recognizing and addressing LDA’s limitations is crucial for effective use in diverse scenarios.
So, this was all about LDA, its mathematics, and its implementation. Hope it was helpful.
Thanks for reading.
FAQs
LDA is a supervised dimensionality reduction technique for classification and feature selection, while PCA is an unsupervised technique for exploratory data analysis and preprocessing.
Merits:
Effective dimensionality reduction
Computationally efficient
Provides insights into data structure
Demerits:
Potential information loss
Sensitivity to outliers
Linearity assumption
In real life, Linear Discriminant Analysis (LDA) is used in medical diagnoses. For example, doctors may analyze MRI or CT scan features to differentiate between different types of cancer. LDA helps identify patterns in these features to classify scans accurately, aiding doctors in diagnosing patients and deciding on treatment plans.
1.Dimensionality reduction
2.Feature extraction
3.Improved classification accuracy
4.Robust performance with small sample sizes
5.Regularization to prevent overfitting
6.Interpretability of results.
Source: An Introduction to Statistical Learning with Applications in R – Gareth James, Daniela









In equation number 8, summation l=1 to K is used whereas l is not used in the terms of summation. Can you please explain?
Linear Discriminant Analysis is a powerful tool for analyzing data. It can be used to identify which variables are most important in predicting outcomes.