4 Ways to Evaluate your Machine Learning Model: Cross-Validation Techniques (with Python code)
This article was published as a part of the Data Science Blogathon
Introduction
Whenever we build any machine learning model, we feed it with initial data to train the model. And then we feed some unknown data (test data) to understand how well the model performs and generalized over unseen data. If the model performs well on the unseen data, it’s consistent and is able to predict with good accuracy on a wide range of input data; then this model is stable.
But this is not the case always! Machine learning models are not always stable and we have to evaluate the stability of the machine learning model. That is where Cross Validation comes into the picture.
“In simple terms, Cross-Validation is a technique used to assess how well our Machine learning models perform on unseen data”
According to Wikipedia, Cross-Validation is the process of assessing how the results of a statistical analysis will generalize to an independent data set.
There are many ways to perform Cross-Validation and we will learn about 4 methods in this article.
Let’s first understand the need for Cross-Validation!
Why do we need Cross-Validation?

Suppose you build a machine learning model to solve a problem, and you have trained the model on a given dataset. When you check the accuracy of the model on the training data, it is close to 95%. Does this mean that your model has trained very well, and it is the best model because of the high accuracy?
No, it’s not! Because your model is trained on the given data, it knows the data well, captured even the minute variations(noise), and has generalized very well over the given data. If you expose the model to completely new, unseen data, it might not predict with the same accuracy and it might fail to generalize over the new data. This problem is called over-fitting.
Sometimes the model doesn’t train well on the training set as it’s not able to find patterns. In this case, it wouldn’t perform well on the test set as well. This problem is called Under-fitting.
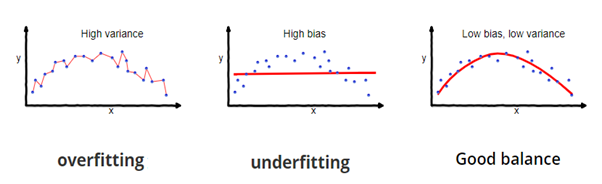
Image Source: fireblazeaischool.in
To overcome over-fitting problems, we use a technique called Cross-Validation.
What is Cross Validation?
Cross-Validation is a resampling technique with the fundamental idea of splitting the dataset into 2 parts- training data and test data. Train data is used to train the model and the unseen test data is used for prediction. If the model performs well over the test data and gives good accuracy, it means the model hasn’t overfitted the training data and can be used for prediction.
Let’s dive deep and learn about some of the model evaluation techniques.
1. Hold Out method
This is the simplest evaluation method and is widely used in Machine Learning projects. Here the entire dataset(population) is divided into 2 sets – train set and test set. The data can be divided into 70-30 or 60-40, 75-25 or 80-20, or even 50-50 depending on the use case. As a rule, the proportion of training data has to be larger than the test data.
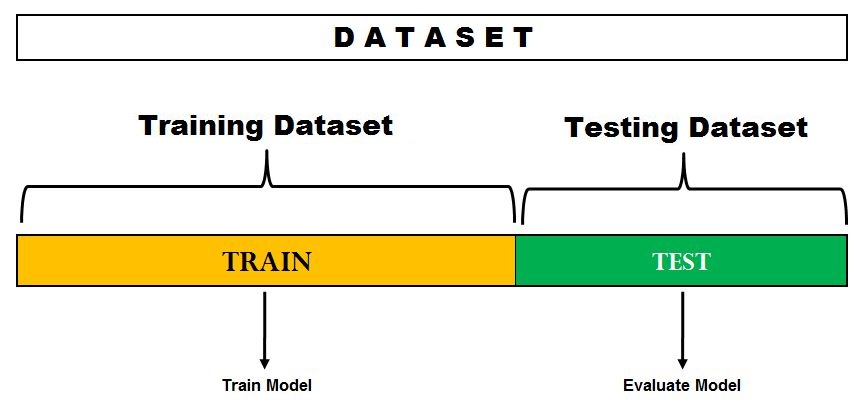
Image Source: DataVedas
The data split happens randomly, and we can’t be sure which data ends up in the train and test bucket during the split unless we specify random_state. This can lead to extremely high variance and every time, the split changes, the accuracy will also change.
There are some drawbacks to this method:
- In the Hold out method, the test error rates are highly variable (high variance) and it totally depends on which observations end up in the training set and test set
- Only a part of the data is used to train the model (high bias) which is not a very good idea when data is not huge and this will lead to overestimation of test error.
One of the major advantages of this method is that it is computationally inexpensive compared to other cross-validation techniques.
Quick implementation of Hold Out method in Python
Python Code:
Output
Train: [50, 10, 40, 20, 80, 90, 60] Test: [30, 100, 70]
Here, random_state is the seed used for reproducibility.
2. Leave One Out Cross-Validation
In this method, we divide the data into train and test sets – but with a twist. Instead of dividing the data into 2 subsets, we select a single observation as test data, and everything else is labeled as training data and the model is trained. Now the 2nd observation is selected as test data and the model is trained on the remaining data.
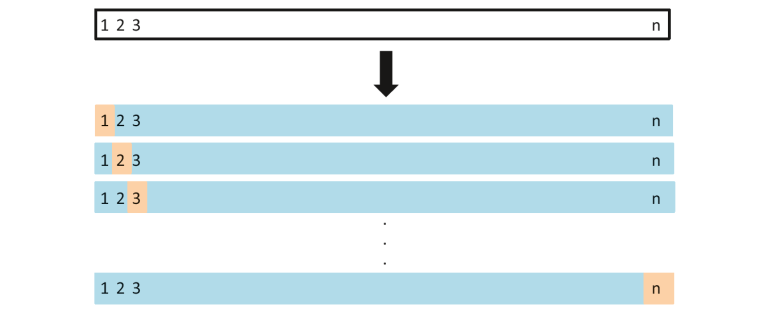
Image Source: ISLR
This process continues ‘n’ times and the average of all these iterations is calculated and estimated as the test set error.
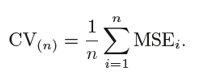
When it comes to test-error estimates, LOOCV gives unbiased estimates (low bias). But bias is not the only matter of concern in estimation problems. We should also consider variance.
LOOCV has an extremely high variance because we are averaging the output of n-models which are fitted on an almost identical set of observations, and their outputs are highly positively correlated with each other.
And you can clearly see this is computationally expensive as the model is run ‘n’ times to test every observation in the data. Our next method will tackle this problem and give us a good balance between bias and variance.
Quick implementation of Leave One Out Cross-Validation in Python
from sklearn.model_selection import LeaveOneOut
X = [10,20,30,40,50,60,70,80,90,100]
l = LeaveOneOut()
for train, test in l.split(X):
print("%s %s"% (train,test))
Output
[1 2 3 4 5 6 7 8 9] [0] [0 2 3 4 5 6 7 8 9] [1] [0 1 3 4 5 6 7 8 9] [2] [0 1 2 4 5 6 7 8 9] [3] [0 1 2 3 5 6 7 8 9] [4] [0 1 2 3 4 6 7 8 9] [5] [0 1 2 3 4 5 7 8 9] [6] [0 1 2 3 4 5 6 8 9] [7] [0 1 2 3 4 5 6 7 9] [8] [0 1 2 3 4 5 6 7 8] [9]
This output clearly shows how LOOCV keeps one observation aside as test data and all the other observations go to train data.
3. K-Fold Cross-Validation
In this resampling technique, the whole data is divided into k sets of almost equal sizes. The first set is selected as the test set and the model is trained on the remaining k-1 sets. The test error rate is then calculated after fitting the model to the test data.
In the second iteration, the 2nd set is selected as a test set and the remaining k-1 sets are used to train the data and the error is calculated. This process continues for all the k sets.
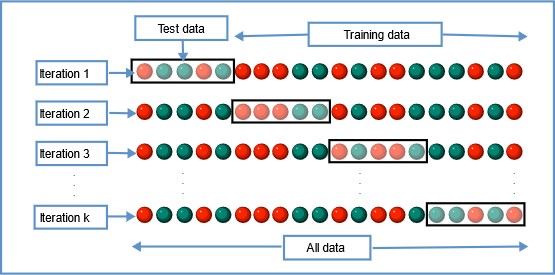
Image Source: Wikipedia
The mean of errors from all the iterations is calculated as the CV test error estimate.
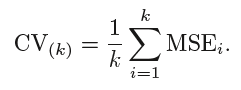
In K-Fold CV, the no of folds k is less than the number of observations in the data (k<n) and we are averaging the outputs of k fitted models that are somewhat less correlated with each other since the overlap between the training sets in each model is smaller. This leads to low variance then LOOCV.
The best part about this method is each data point gets to be in the test set exactly once and gets to be part of the training set k-1 times. As the number of folds k increases, the variance also decreases (low variance). This method leads to intermediate bias because each training set contains fewer observations (k-1)n/k than the Leave One Out method but more than the Hold Out method.
Typically, K-fold Cross Validation is performed using k=5 or k=10 as these values have been empirically shown to yield test error estimates that neither have high bias nor high variance.
The major disadvantage of this method is that the model has to be run from scratch k-times and is computationally expensive than the Hold Out method but better than the Leave One Out method.
Simple implementation of K-Fold Cross-Validation in Python
from sklearn.model_selection import KFold
X = ["a",'b','c','d','e','f']
kf = KFold(n_splits=3, shuffle=False, random_state=None)
for train, test in kf.split(X):
print("Train data",train,"Test data",test)
Output
Train: [2 3 4 5] Test: [0 1] Train: [0 1 4 5] Test: [2 3] Train: [0 1 2 3] Test: [4 5]
4. Stratified K-Fold Cross-Validation
This is a slight variation from K-Fold Cross Validation, which uses ‘stratified sampling’ instead of ‘random sampling.’
Let’s quickly understand what stratified sampling is and how is it different from random sampling.
Suppose your data contains reviews for a cosmetic product used by both the male and female population. When we perform random sampling to split the data into train and test sets, there is a possibility that most of the data representing males is not represented in training data but might end up in test data. When we train the model on sample training data that is not a correct representation of the actual population, the model will not predict the test data with good accuracy.
This is where Stratified Sampling comes to the rescue. Here the data is split in such a way that it represents all the classes from the population.
Let’s consider the above example which has a cosmetic product review of 1000 customers out of which 60% is female and 40% is male. I want to split the data into train and test data in proportion (80:20). 80% of 1000 customers will be 800 which will be chosen in such a way that there are 480 reviews associated with the female population and 320 representing the male population. In a similar fashion, 20% of 1000 customers will be chosen for the test data ( with the same female and male representation).
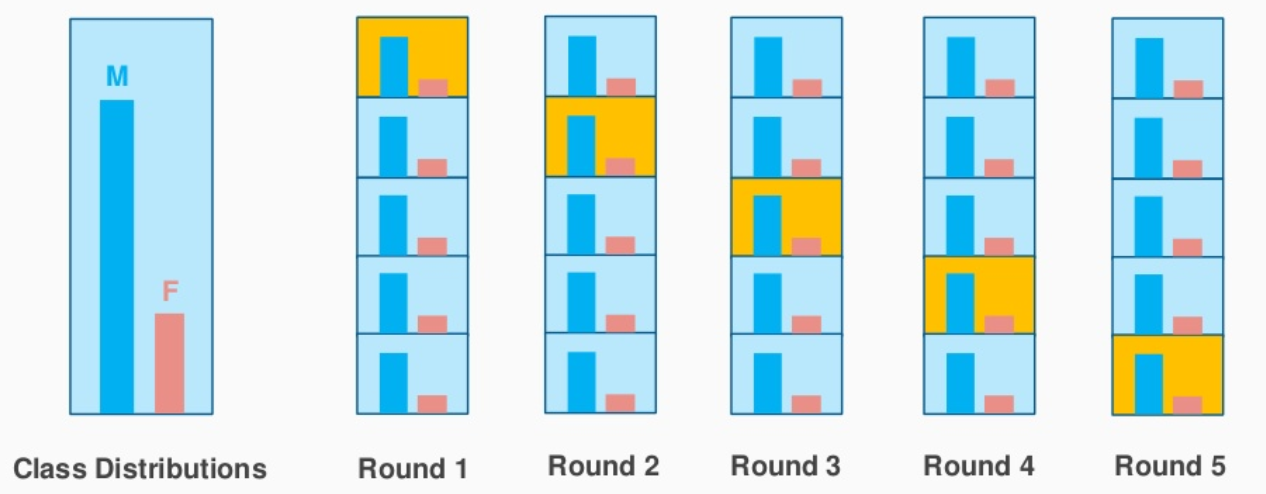
Image Source: stackexchange.com
This is exactly what stratified K-Fold CV does and it will create K-Folds by preserving the percentage of sample for each class. This solves the problem of random sampling associated with Hold out and K-Fold methods.
Quick implementation of Stratified K-Fold Cross-Validation in Python
from sklearn.model_selection import StratifiedKFold
X = np.array([[1,2],[3,4],[5,6],[7,8],[9,10],[11,12]])
y= np.array([0,0,1,0,1,1])
skf = StratifiedKFold(n_splits=3,random_state=None,shuffle=False)
for train_index,test_index in skf.split(X,y):
print("Train:",train_index,'Test:',test_index)
X_train,X_test = X[train_index], X[test_index]
y_train,y_test = y[train_index], y[test_index]
Output
Train: [1 3 4 5] Test: [0 2] Train: [0 2 3 5] Test: [1 4] Train: [0 1 2 4] Test: [3 5]
The output clearly shows the stratified split done based on the classes ‘0’ and ‘1’ in ‘y’.
Bias – Variance Tradeoff
When we consider the test error rate estimates, K-Fold Cross Validation gives more accurate estimates than Leave One Out Cross-Validation. Whereas Hold One Out CV method usually leads to overestimates of the test error rate, because in this approach, only a portion of the data is used to train the machine learning model.
When it comes to bias, the Leave One Out Method gives unbiased estimates because each training set contains n-1 observations (which is pretty much all of the data). K-Fold CV leads to an intermediate level of bias depending on the number of k-folds when compared to LOOCV but it’s much lower when compared to the Hold Out Method.
To conclude, the Cross-Validation technique that we choose highly depends on the use case and bias-variance trade-off.
If you have read this article this far, here is a quick bonus for you 👏
sklearn.model_selection has a method cross_val_score which simplifies the process of cross-validation. Instead of iterating through the complete data using the ‘split’ function, we can use cross_val_score and check the accuracy score for the chosen cross-validation method
You can check out my Github for python implementation of different cross-validation methods on the UCI Breast Cancer data from Kaggle.
Frequently Asked Questions
A. Cross-validation is a technique used in machine learning and statistical modeling to assess the performance of a model and to prevent overfitting. It involves dividing the dataset into multiple subsets, using some for training the model and the rest for testing, multiple times to obtain reliable performance metrics.
Types of Cross-validation:
1. K-Fold Cross-Validation: The data is divided into K subsets (or “folds”). The model is trained K times, using K-1 folds for training and one fold for testing in each iteration.
2. Leave-One-Out Cross-Validation (LOOCV): K-Fold CV with K equal to the number of data points, i.e., each data point is used once as a test set, and the model is trained K times.
3. Stratified K-Fold Cross-Validation: It ensures that the class distribution remains similar in each fold, important when dealing with imbalanced datasets.
4. Time Series Cross-Validation: For time-dependent data, it uses a series of temporally ordered training and testing sets, preventing the use of future data for training.
5. Shuffle-Split Cross-Validation: Randomly shuffles the data, and then splits it into training and testing sets multiple times.
6. Group K-Fold Cross-Validation: Useful when the data contains groups, like multiple samples from the same subject, and ensures all samples from a group are kept together in the same fold.
Cross-validation provides a more robust assessment of a model’s performance and helps in selecting hyperparameters, assessing generalization, and avoiding overfitting by evaluating the model on different data points than those used for training.
A. K-fold cross-validation is used to assess the performance and generalization of a machine learning model. It divides the dataset into K subsets, trains the model K times using K-1 folds for training and one fold for testing in each iteration. This method helps in obtaining more reliable performance metrics, choosing hyperparameters, and preventing overfitting by evaluating the model on different data subsets.
Below are some of my articles on Machine Learning
A Comprehensive Guide to Data Analysis using Pandas
If you would like to share your thoughts, you can connect with me on LinkedIn.
Happy Learning!
The media shown in this article are not owned by Analytics Vidhya and is used at the Author’s discretion.









I would like to point out a discrepancy in the article. Under LOOCV, following paragraph is present : -------------------- "LOOCV has an extremely high variance because we are averaging the output of n-models which are fitted on an almost identical set of observations, and their outputs are highly positively correlated with each other." -------------------- Under K-Fold CV following line is present: -------------------- As the number of folds k increases, the variance also decreases (low variance). -------------------- But as we increase the number of folds, don't you think the variance should increase? The highest variance one might get is in LOOCV where one sample is left out, as the number of "k" folds increases, the sample per fold decreases and hence variance goes higher. Please shed some light on this.