Introduction
Machine learning algorithms are classified into three types: supervised learning, unsupervised learning, and reinforcement learning. K–means clustering is an unsupervised machine learning technique. When the output or response variable is not provided, this algorithm is used to categorize the data into distinct clusters for getting a better understanding of it. It is also known as a data-driven machine learning approach since it clusters data based on hidden patterns, insights, and similarities in the data.
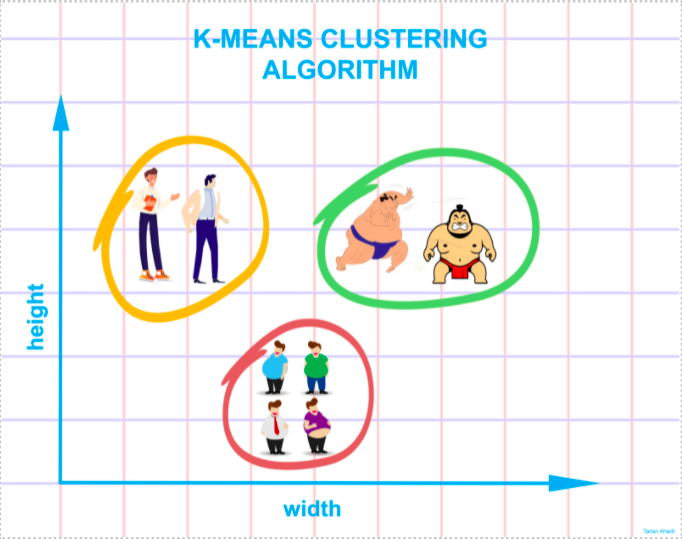
Consider the following diagram: if you are asked to group the people in the picture into different clusters or groups and you know nothing about them, you will undoubtedly try to locate the qualities, features, or physical attributes that these people are sharing. After observing these people, you conclude that they can be segregated based on their height and width; given that you have no prior knowledge about these people. The k-means clustering executes a roughly equivalent job. It attempts to categorize data into clusters based on similarities and hidden patterns. ‘K” in K-means clustering refers to the number of clusters, the algorithm will generate in the data.
K–Means Clustering: How it Works?
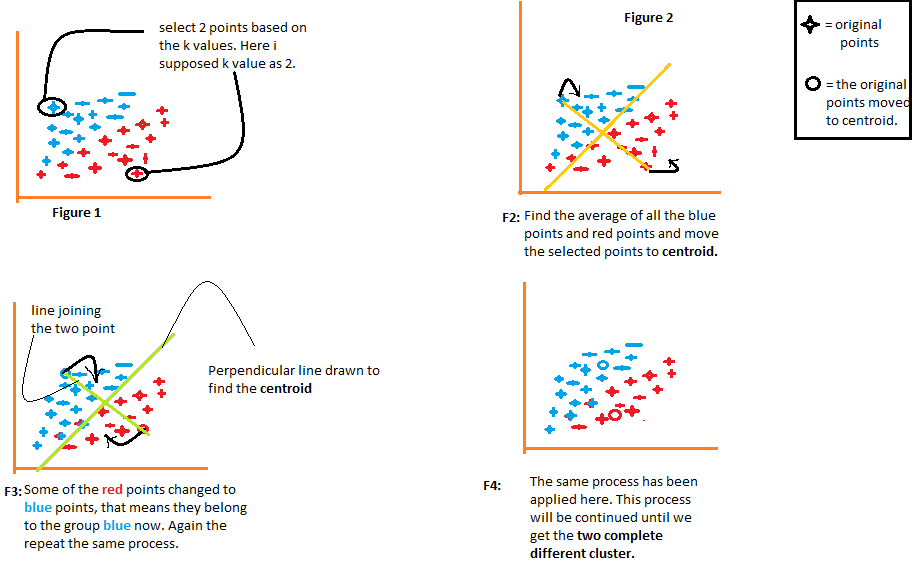
1) The algorithm arbitrarily chooses the k number of centroids, as indicated in figure 1 of the following diagram. Where k is the number of clusters the algorithm would create. Let’s say we want the algorithm to create two clusters from the data, so we’ll set the value of k to 2.
2) It then clusters the data into two parts using the distances calculated from both the centroids, as illustrated in Figure 2. Each point’s distance from both centroids is computed individually and will subsequently be added to the group of that centroid with which the distance is shortest.
The algorithm also draws a line joining both the centroids and a perpendicular line to it that attempts to cluster the data into two groups.
3) Once all of the data points are grouped based on their minimal distances from the corresponding centroids, the algorithm then calculates the mean of each group. The mean and centroid values of each group are then compared. If the centroid value differs from the mean, then the centroid is shifted to the group’s mean value. Both the “Red” and “Blue” centroids are relocated to the group’s mean in figure 3 of the following diagram.
It groups the data once again using these updated centroids. Because of the shift in the centroids’ positions, some data points may now get shifted in the other group.
4) Again, it calculates the mean and compares it to the centroid for newly generated groups. If both of them are different, the centroid will again be relocated to the group mean. This process of computing the mean and comparing it to the centroid is iterated till the centroid and mean values become equal (centroid value = group mean). This is the point at which the algorithm has segmented the data into ‘K’ clusters (2 in this case).
How to figure out what the optimal value of k is?
The first step is to provide a value for k. Every subsequent step executed by the algorithm is fully dependent on the value of k specified. This k value assists the algorithm in determining the number of clusters that should be generated. This emphasizes the importance of providing the precise value for k. Here, a method known as the “Elbow Method” is used to determine the correct value of k. This is a graph of ‘Number of clusters K’ vs “Total Within Sum of Square”. Discrete values of k are plotted on the x-axis, while cluster sums of squares are plotted on the y-axis.
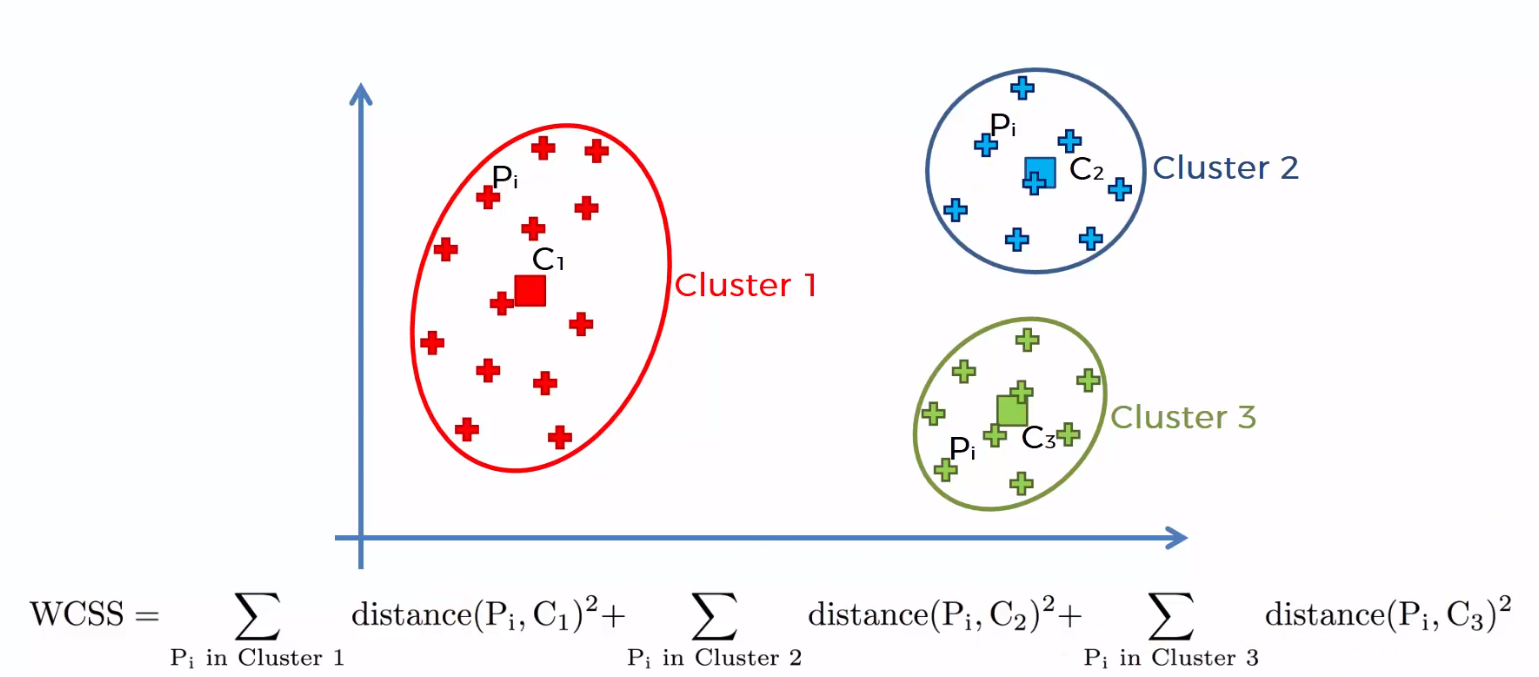
The summation of squared distances between individual points and centroid in each group, followed by the summing of squared distances for all clusters, is referred to as the “Within Cluster Sum of Square”. You will be able to understand this with the help of the following steps.
1) Calculate the distance between the centroid and each point in the cluster, square it, then sum the squared distances for all of the points in the cluster.
2) Find the sum of squared distances for the remaining clusters in the same way.
3) Finally, add all of the cluster summations together to get the value of the “Within Cluster Sum of Square” as shown in the following figure.
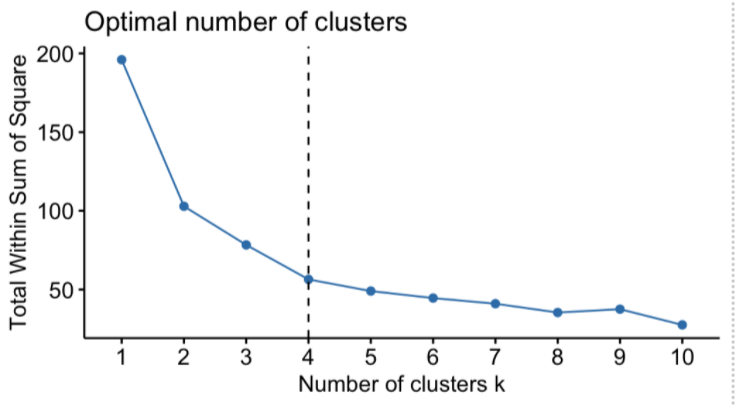
The “total within the sum of square” starts to decrease as the value of k increases. The plot between the number of clusters and the total within the sum of squares is shown in the figure below. The optimal number of clusters, or the correct value of k, is the point at which the value begins to decrease slowly; this is known as the ‘elbow point’, and the elbow point in the following plot is k = 4. The “Elbow Method” is named for the plot’s resemblance to the elbow, and the optimal point for “k” is the elbow point.
Advantages of k-Means Clustering
1) The labeled data isn’t required. Since so much real-world data is unlabeled, as a result, it is frequently utilized in a variety of real-world problem statements.
2) It is easy to implement.
3) It can handle massive amounts of data.
4) When the data is large, it works faster than hierarchical clustering (for small k).
Disadvantages of K-Means Clustering
1) K value is required to be selected manually using the “elbow method”.
2) The presence of outliers would have an adverse impact on the clustering. As a result, outliers must be eliminated before using k-means clustering.
3) Clusters do not cross across; a point may only belong to one cluster at a time. As a result of the lack of overlapping, certain points are placed in the incorrect clusters.
K-Means Clustering with R
- We will import the following libraries into our work.
library(caret)
library(ggplot2)
library(dplyr)
- We’ll be working with the iris data, which contains three classes: “Iris-setosa,” “Iris-versicolor,” and “Iris-virginica”.
data <- read.csv(“iris.csv”, header =T)
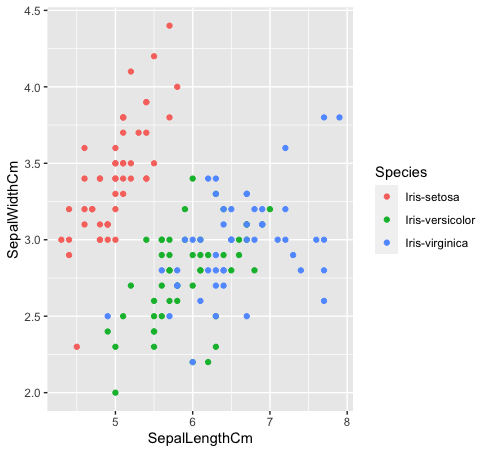
- Let’s look at how these three classes are related to one another. The species “Iris-versicolor” (green) and “Iris-verginica” (blue) are not linearly separable. As can be seen in the plot below, they are intermixing.
data %>% ggplot(aes(SepalLengthCm, SepalWidthCm, color= Species))+
geom_point()
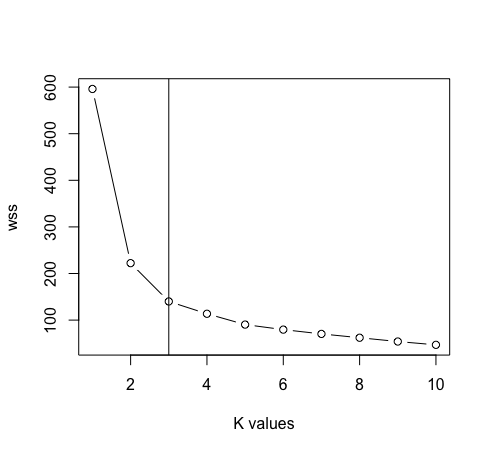
- After removing the species column from the data. We’ll now use the elbow method’s plot between “Within cluster sum of square” and “Values of k” to determine the suitable value of k. K=3 is the best value for k in this case (Note: There are 3 classes in the original iris data, which ensures the accuracy of the value of k).
data <- data[, -5] maximum <- 10
scal <- scale(data)
wss <- sapply(1:maximum, function(k){kmeans(scal, k, nstart = 50, iter.max = 15)$tot.withinss})
plot(1:max, wss,type=”b”, xlab= “k values”)
abline(v=3)
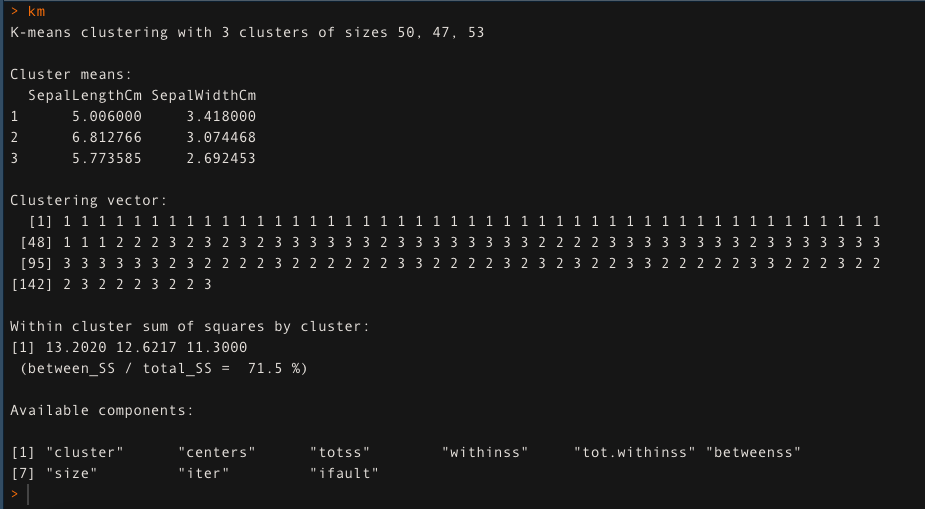
- For k=3, apply the K-means clustering algorithm. The K-means clustering approach is explaining 71.5 % of the data variability in this case.
km <- kmeans(data[,1:2], k=3, iter.max = 50)
km
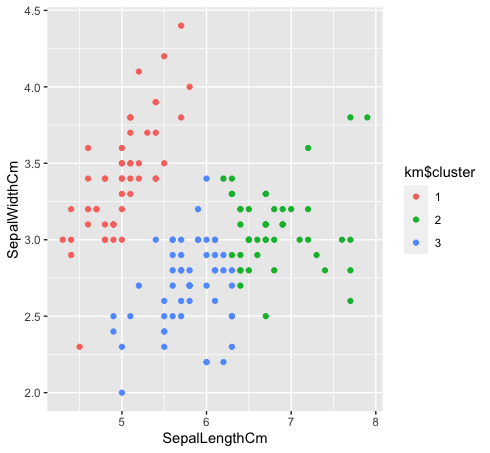
- Let’s look at how the three classes are clustered using k-means clustering. K-means clustering won’t create overlapping clusters, as we all know. Since “green” and “blue” species are not linearly separable in the original data, k-means clustering could not capture it because of having tight clusters.
km$cluster <- as.factor(km$cluster)
data %>% ggplot(aes(SepalLengthCm, SepalWidthCm, color= km$cluster))+
geom_point()
An Article by~
Shivam Sharma.
Phone: +91 7974280920
E-Mail: [email protected]
The media shown in this article on K-Means Clustering Algorithm are not owned by Analytics Vidhya and is used at the Author’s discretion.