Deep Learning based Malaria Detection Model for Beginners
This article was published as a part of the Data Science Blogathon
Overview
Malaria is a significant burden on our healthcare system and it is the major cause of death in many developing countries. It is endemic in some parts of the world which means that the disease is regularly found in the region. Therefore, early testing is necessary to detect malaria and save lives. Thus, it gives us the motivation to make malaria diagnosis more effective and faster. A specialized technology proves essential to combat this problem.
For Malaria diagnosis, the RDT and microscopic diagnosis are the most used clinical methods. RDT is an effective and faster tool. Also, it does not require the presence of a trained medical professional. But RDT has few drawbacks like susceptibility to damage by heat and humidity and higher cost compared to a light microscope. The microscopic diagnosis systems do not suffer from these shortcomings, but it requires the presence of a trained microscopist. Machine learning has recently made headlines in the medical field. The machine learning approaches have proved to be successful in the diagnosis of a disease.
Problem definition for Deep Learning-based Malaria Detection
The main aim of this malaria detection system is to address the challenges in the existing system by automating the process of malaria detection using Machine learning and image processing.
Demo video of the Malaria Detection Model
Dataset to be used for Malaria Detection
The dataset used for this system is provided by ‘National Institutes of Health’, which consists of 30000 cell images. The dataset can be downloaded from here.
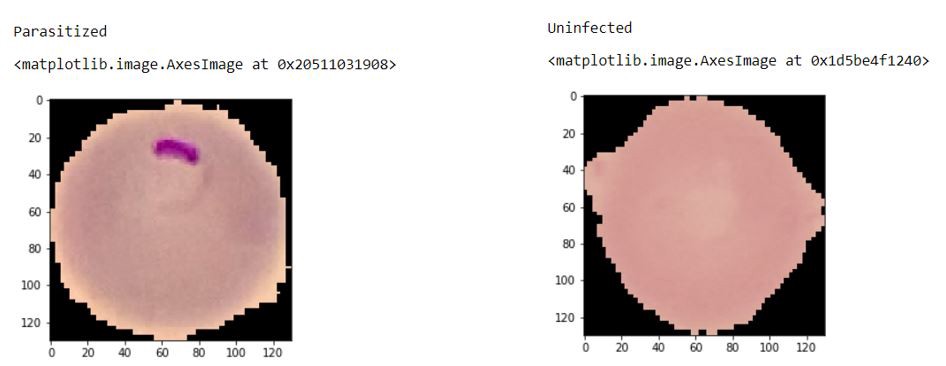
This dataset consists of two folders: the test and train folder with the testing and training images, respectively. Further, there are two more folders -parasitized folder with the malaria-infected cell images and the uninfected folder with the normal cell images. As we have an equal distribution of cell images, so we don’t have to deal with the problems of imbalanced data, and so it has fewer chances of being biased towards a particular class.
Algorithm for Malaria Detection
The input image is first processed to remove unwanted noise from the RGB cell image. The preprocessed image is then given as an input to the segmentation stage. The image is segmented to extract the Region of interest from the image, and we get the segmented image. We then feed the image as an input to the feature extraction stage, where the output will be the feature vectors. The next stage is the classification stage, where the input will be the feature vectors, and output is the classified label as parasitic and non-parasitic.
Image preprocessing
This is the first stage. The aim of data preprocessing is to clean the cell images. This stage will remove the unwanted noise
kernel1 = np.ones((9,9),np.uint8) clean1 = cv2.morphologyEx(image, cv2.MORPH_OPEN, kernel1)
Image segmentation
This is the second stage where the image is segmented to extract the region of interest from the image. Here we will be using two image segmentation techniques as follows:
Otsu’s segmentation
It is used to perform automatic segmentation by image thresholding of the cell images. It returns a single intensity threshold value which separates pixels into two classes.
# SEGMENTATION
import numpy as np
import cv2
from matplotlib import pyplot as plt
img = cv2.imread(r'C33P1thinF_IMG_20150619_114756a_cell_181.png')
b,g,r = cv2.split(img)
rgb_img = cv2.merge([r,g,b])
gray = cv2.cvtColor(img,cv2.COLOR_BGR2GRAY)
ret, thresh = cv2.threshold(gray,0,255,cv2.THRESH_BINARY_INV+cv2.THRESH_OTSU)
# noise removal
kernel = np.ones((2,2),np.uint8)
#opening = cv2.morphologyEx(thresh,cv2.MORPH_OPEN,kernel, iterations = 2)
closing = cv2.morphologyEx(thresh,cv2.MORPH_CLOSE,kernel, iterations = 2)
# sure background area
sure_bg = cv2.dilate(closing,kernel,iterations=3)
# Finding sure foreground area
dist_transform = cv2.distanceTransform(sure_bg,cv2.DIST_L2,3)
# Threshold
ret, sure_fg = cv2.threshold(dist_transform,0.1*dist_transform.max(),255,0)
# Finding unknown region
sure_fg = np.uint8(sure_fg)
unknown = cv2.subtract(sure_bg,sure_fg)
# Marker labelling
ret, markers = cv2.connectedComponents(sure_fg)
# Add one to all labels so that sure background is not 0, but 1
markers = markers+1
# Now, mark the region of unknown with zero
markers[unknown==255] = 0
markers = cv2.watershed(img,markers)
img[markers == -1] = [255,0,0]
plt.subplot(211),plt.imshow(rgb_img)
plt.title('Input Image'), plt.xticks([]), plt.yticks([])
plt.subplot(212),plt.imshow(thresh, 'gray')
plt.imsave(r'thresh.png',thresh)
plt.title("Otsu's binary threshold"), plt.xticks([]), plt.yticks([])
plt.tight_layout()
plt.show()
Watershed segmentation
Watershed segmentation is an image segmentation algorithm that segments an image into several catchment basins or regions. The input image can be segmented into regions where conceptually rainwater would flow into the same lake and this will result in partitioning the image into catchment basins and watershed lines.
import numpy as np
import cv2
from matplotlib import pyplot as plt
img = cv2.imread(r'C33P1thinF_IMG_20150619_114756a_cell_181.png')
b,g,r = cv2.split(img)
rgb_img = cv2.merge([r,g,b])
gray = cv2.cvtColor(img,cv2.COLOR_BGR2GRAY)
ret, thresh = cv2.threshold(gray,0,255,cv2.THRESH_BINARY_INV+cv2.THRESH_OTSU)
plt.subplot(211),plt.imshow(closing, 'gray')
plt.title("morphologyEx:Closing:2x2"), plt.xticks([]), plt.yticks([])
plt.subplot(212),plt.imshow(sure_bg, 'gray')
plt.imsave(r'dilation.png',sure_bg)
plt.title("Dilation"), plt.xticks([]), plt.yticks([])
plt.tight_layout()
plt.show()
Feature extraction & Classification
The segmented image is given as an input and we get the feature vectors which are then used for the classification task. Here we will be using Convolutional Neural Network (CNN) model with the Relu activation function.
Convolutional Neural Network
CNN is a model which is designed to process arrays of data such as images. The first step here will be to resize all images as CNN cannot train images of different sizes. We compute the mean for both the dimensions and resize all the images. Now we can use random transform and we will double our dataset.
image_gen = ImageDataGenerator(rotation_range=20,
width_shift_range=0.10,
height_shift_range=0.10,
rescale=1/255,
shear_range=0.1,
zoom_range=0.1,
horizontal_flip=True,
fill_mode='nearest'
)
plt.imshow(image_gen.random_transform(parasitic_img))
Build a baseline model for Malaria Detection
model = Sequential()
model.add(Conv2D(filters=32, kernel_size=(3,3),input_shape=image_shape, activation='relu',))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(filters=64, kernel_size=(3,3),input_shape=image_shape, activation='relu',))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(filters=64, kernel_size=(3,3),input_shape=image_shape, activation='relu',))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(128))
model.add(Activation('relu'))
# Dropouts help reduce overfitting by randomly turning neurons off during training.
# Here we say randomly turn off 50% of neurons.
model.add(Dropout(0.5))
# Last layer, remember its binary so we use sigmoid
model.add(Dense(1))
model.add(Activation('sigmoid'))
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
The sequential model is used here. In the first layer which is the Convolution layer, we will place the filter on top of the input matrix and then compute the value and will be doing a stride jump of 1. This extracts features from the image. Also, we can use Padding if the filter does not fit perfectly fit the input image. Here we will be using the Relu activation function.
ReLU(x)=max(0,x)
Max pooling selects the maximum element and extracts the most prominent features from the image. Last is the fully connected layer, where the input to the fully connected layer will be the output from the Max Pooling Layer; it is flattened and then fed into the fully connected layer.
Results of the Malaria Detection Model
Performance of this system is evaluated using parameters like Accuracy, recall, precision, and F1-score. This model achieved an accuracy of 94% and 93% precision.
Graphical User Interface
The GUI for this system is created using Python’s Tkinter library as follows:
from tkinter import *
from tkinter import ttk
from tkinter import filedialog
class Root(Tk):
def __init__(self):
super(Root, self).__init__()
self.title("Python Tkinter Dialog Widget")
self.minsize(640, 400)
self.labelFrame = ttk.LabelFrame(self, text = "Open File")
self.labelFrame.grid(column = 0, row = 1, padx = 20, pady = 20)
self.button()
self.button1()
def button(self):
self.button = ttk.Button(self.labelFrame, text = "Browse A File",command = self.fileDialog)
self.button.grid(column = 1, row = 1)
def fileDialog(self):
self.filename = filedialog.askopenfilename(initialdir = "/", title = "Select A File", filetype =
(("png files","*.png"),("all files","*.*")) )
self.label = ttk.Label(self.labelFrame, text = "")
self.label.grid(column = 1, row = 2)
self.label.configure(text = self.filename)
def button1(self):
self.button = ttk.Button(self.labelFrame, text = "submit", command = self.get_prediction)
self.button.grid(column = 1, row = 20)
def get_prediction(self):
s=model.predict(my_image)
if(s==[[1.]]):
self.label.configure(text="non parasitic")
else:
self.label.configure(text="parasitic")
root = Root()
root.mainloop()
The output of the model:
Conclusion
This model achieved an accuracy of 94%. Using data augmentation in the Convolutional Neural Network approach decreases the chances of overfitting. Thus, Malaria detection systems using deep learning proved to be faster than most of the traditional techniques. The system is easy to use and user-friendly.








