If you are here, you are already introduced to the concept of logistic regression and probably have had your hands dirty working on different datasets. The scikitlearn’s LogisticRegression is by far the best tool to use for any logistic regression task, but it is a good exercise to fiddle around and write your logistic regression algorithm and see how your algorithm fares. It always helps a great deal to write algorithms from scratch, provides you with details that you otherwise have missed, It consolidates your knowledge regarding the topic. It will be helpful if you have a prior understanding of matrix algebra and Numpy. In this article, we will only be dealing with Numpy arrays. Well, let’s get started,
This article was published as a part of the Data Science Blogathon.
Table of contents
Import libraries for Logistic Regression
import numpy as np from numpy import log,dot,e,shape import matplotlib.pyplot as plt
For this article, we will be using sklearn’s make_classification dataset with four features:
from sklearn.datasets import make_classification
X,y = make_classification(n_features = 4,n_classes=2)
from sklearn.model_selection import train_test_split
X_tr,X_te,y_tr,y_te = train_test_split(X,y,test_size=0.1)
print(X_tr.shape, X_te.shape)Standardization
def standardize(X_tr):
for i in range(shape(X_tr)[1]):
X_tr[:,i] = (X_tr[:,i] - np.mean(X_tr[:,i]))/np.std(X_tr[:,i])
Initializing Parameters
def initialize(self,X):
weights = np.zeros((shape(X)[1]+1,1))
X = np.c_[np.ones((shape(X)[0],1)),X]
return weights,X
Sigmoid Function
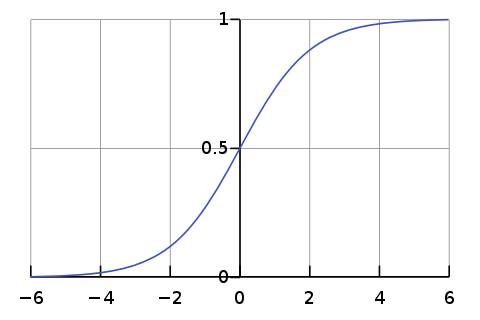
def sigmoid(self,z):
sig = 1/(1+e**(-z))
return sig
Cost Function
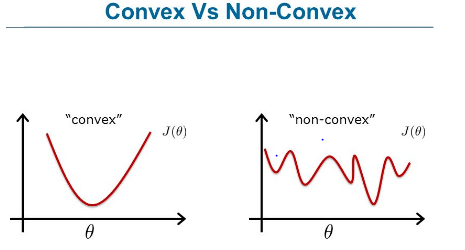
Image source: www.fromthegenesis.com/artificial-neural-network-part-7/
def cost(theta):
z = dot(X,theta)
cost0 = y.T.dot(log(self.sigmoid(z)))
cost1 = (1-y).T.dot(log(1-self.sigmoid(z)))
cost = -((cost1 + cost0))/len(y)
return cost
Gradient Descent
or,
This is the vectorised form of the gradient descent expression, which we will be using in our code.
def fit(self,X,y,alpha=0.001,iter=100):
params,X = self.initialize(X)
cost_list = np.zeros(iter,)
for i in range(iter):
params = params - alpha * dot(X.T, self.sigmoid(dot(X,params)) - np.reshape(y,(len(y),1)))
cost_list[i] = cost(params)
self.params = params
return cost_list
Prediction
def predict(self,X):
z = dot(self.initialize(X)[1],self.weights)
lis = []
for i in self.sigmoid(z):
if i>0.5:
lis.append(1)
else:
lis.append(0)
return lis
F1-Score
recall = TP/(TP+FN)
def f1_score(y,y_hat):
tp,tn,fp,fn = 0,0,0,0
for i in range(len(y)):
if y[i] == 1 and y_hat[i] == 1:
tp += 1
elif y[i] == 1 and y_hat[i] == 0:
fn += 1
elif y[i] == 0 and y_hat[i] == 1:
fp += 1
elif y[i] == 0 and y_hat[i] == 0:
tn += 1
precision = tp/(tp+fp)
recall = tp/(tp+fn)
f1_score = 2*precision*recall/(precision+recall)
return f1_score
Putting Everything Together: Logistic Regression
Now that we are done with every part, we will put everything together in a single class.
import numpy as np
from numpy import log,dot,exp,shape
import matplotlib.pyplot as plt
from sklearn.datasets import make_classification
X,y = make_classification(n_featues=4)
from sklearn.model_selection import train_test_split
X_tr,X_te,y_tr,y_te = train_test_split(X,y,test_size=0.1
def standardize(X_tr):
for i in range(shape(X_tr)[1]):
X_tr[:,i] = (X_tr[:,i] - np.mean(X_tr[:,i]))/np.std(X_tr[:,i])
def F1_score(y,y_hat):
tp,tn,fp,fn = 0,0,0,0
for i in range(len(y)):
if y[i] == 1 and y_hat[i] == 1:
tp += 1
elif y[i] == 1 and y_hat[i] == 0:
fn += 1
elif y[i] == 0 and y_hat[i] == 1:
fp += 1
elif y[i] == 0 and y_hat[i] == 0:
tn += 1
precision = tp/(tp+fp)
recall = tp/(tp+fn)
f1_score = 2*precision*recall/(precision+recall)
return f1_score
class LogidticRegression:
def sigmoid(self,z):
sig = 1/(1+exp(-z))
return sig
def initialize(self,X):
weights = np.zeros((shape(X)[1]+1,1))
X = np.c_[np.ones((shape(X)[0],1)),X]
return weights,X
def fit(self,X,y,alpha=0.001,iter=400):
weights,X = self.initialize(X)
def cost(theta):
z = dot(X,theta)
cost0 = y.T.dot(log(self.sigmoid(z)))
cost1 = (1-y).T.dot(log(1-self.sigmoid(z)))
cost = -((cost1 + cost0))/len(y)
return cost
cost_list = np.zeros(iter,)
for i in range(iter):
weights = weights - alpha*dot(X.T,self.sigmoid(dot(X,weights))-np.reshape(y,(len(y),1)))
cost_list[i] = cost(weights)
self.weights = weights
return cost_list
def predict(self,X):
z = dot(self.initialize(X)[1],self.weights)
lis = []
for i in self.sigmoid(z):
if i>0.5:
lis.append(1)
else:
lis.append(0)
return lis
standardize(X_tr)
standardize(X_te)
obj1 = LogidticRegression()
model= obj1.fit(X_tr,y_tr)
y_pred = obj1.predict(X_te)
y_train = obj1.predict(X_tr)
#Let's see the f1-score for training and testing data
f1_score_tr = F1_score(y_tr,y_train)
f1_score_te = F1_score(y_te,y_pred)
print(f1_score_tr)
print(f1_score_te)
output:0.9777777777777777 0.9090909090909091
Now. let’s see if our cost function is descending or not
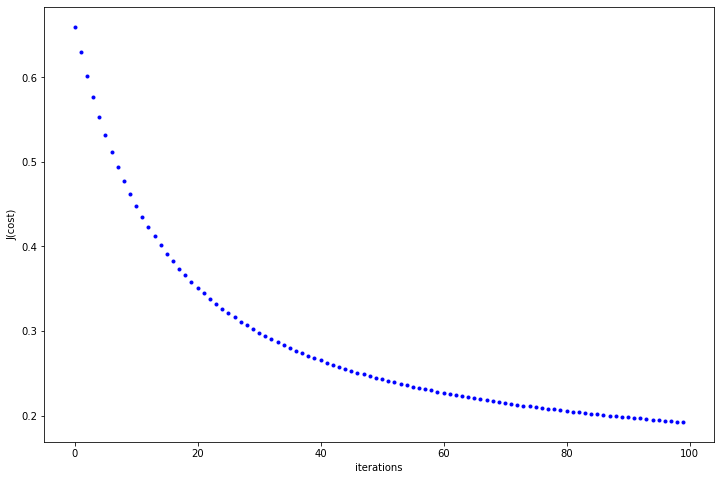
In the above plot, we can see that the cost function decreases with every iteration and almost gets flattened as we move towards 100. You can fiddle around with hyper-parameters and see the behaviour of cost function.
Now, let’s see how our logistic regression fares in comparison to sklearn’s logistic regression.
from sklearn.linear_model import LogisticRegression from sklearn.metrics import f1_score model = LogisticRegression().fit(X_tr,y_tr) y_pred = model.predict(X_te) print(f1_score(y_te,y_pred))
output: 0.9090909090909091
Great!
Logistic Regression EndNote
This article went through different parts of logistic regression and saw how we could implement it through raw python code. But if you are working on some real project, it’s better to opt for Scikitlearn rather than writing it from scratch as it is quite robust to minor inconsistencies and less time-consuming.
Happy coding!
The media shown in this article is not owned by Analytics Vidhya and are used at the Author’s discretion.







Line 7 you have to close the bracket :)