A Complete Tutorial to learn Data Science in R from Scratch
Overview
- Free tutorial to learn Data Science in R for beginners
- Covers predictive modeling, data manipulation, data exploration, and machine learning algorithms in R
Introduction
R is a powerful language used widely for data analysis and statistical computing. It was developed in early 90s. Since then, endless efforts have been made to improve R’s user interface. The journey of R language from a rudimentary text editor to interactive R Studio and more recently Jupyter Notebooks has engaged many data science communities across the world.
This was possible only because of generous contributions by R users globally. Inclusion of powerful packages in R has made it more and more powerful with time. Packages such as dplyr, tidyr, readr, data.table, SparkR, ggplot2 have made data manipulation, visualization and computation much faster.
But, what about Machine Learning?
My first impression of R was that it’s just a software for statistical computing. Good thing, I was wrong! R has enough provisions to implement machine learning algorithms in a fast and simple manner.
This is a complete tutorial to learn data science and machine learning using R. By the end of this tutorial, you will have a good exposure to building predictive models using machine learning on your own.
Note: No prior knowledge of data science / analytics is required. However, prior knowledge of algebra and statistics will be helpful.

A Complete Tutorial to learn Data Science in R from Scratch
Table of Contents
- Basics of R Programming for Data Science
- Why learn R ?
- How to install R / R Studio ?
- How to install R packages ?
- Basic computations in R
- Essentials of R Programming
- Data Types and Objects in R
- Control Structures (Functions) in R
- Useful R Packages
- Exploratory Data Analysis in R
- Basic Graphs
- Treating Missing values
- Working with Continuous and Categorical Variables
- Data Manipulation in R
- Feature Engineering
- Label Encoding / One Hot Encoding
- Predictive Modeling using Machine Learning in R
- Linear Regression
- Decision Tree
- Random Forest
Note: The data set used in this article is from Big Mart Sales Prediction.
1. Basics of R Programming
Why learn R ?
I don’t know if I have a solid reason to convince you, but let me share what got me started. I have no prior coding experience. Actually, I never had computer science in my subjects. I came to know that to learn data science, one must learn either R or Python as a starter. I chose the former. Here are some benefits I found after using R:
- The style of coding is quite easy.
- It’s open source. No need to pay any subscription charges.
- Availability of instant access to over 7800 packages customized for various computation tasks.
- The community support is overwhelming. There are numerous forums to help you out.
- Get high performance computing experience ( require packages)
- One of highly sought skill by analytics and data science companies.
There are many more benefits. But, these are the ones which have kept me going. If you think they are exciting, stick around and move to next section. And, if you aren’t convinced, you may like Complete Python Tutorial from Scratch.
How to install R / R Studio ?
You could download and install the old version of R. But, I’d insist you to start with RStudio. It provides much better coding experience. For Windows users, R Studio is available for Windows Vista and above versions. Follow the steps below for installing R Studio:
- Go to https://www.rstudio.com/products/rstudio/download/
- In ‘Installers for Supported Platforms’ section, choose and click the R Studio installer based on your operating system. The download should begin as soon as you click.
- Click Next..Next..Finish.
- Download Complete.
- To Start R Studio, click on its desktop icon or use ‘search windows’ to access the program. It looks like this:
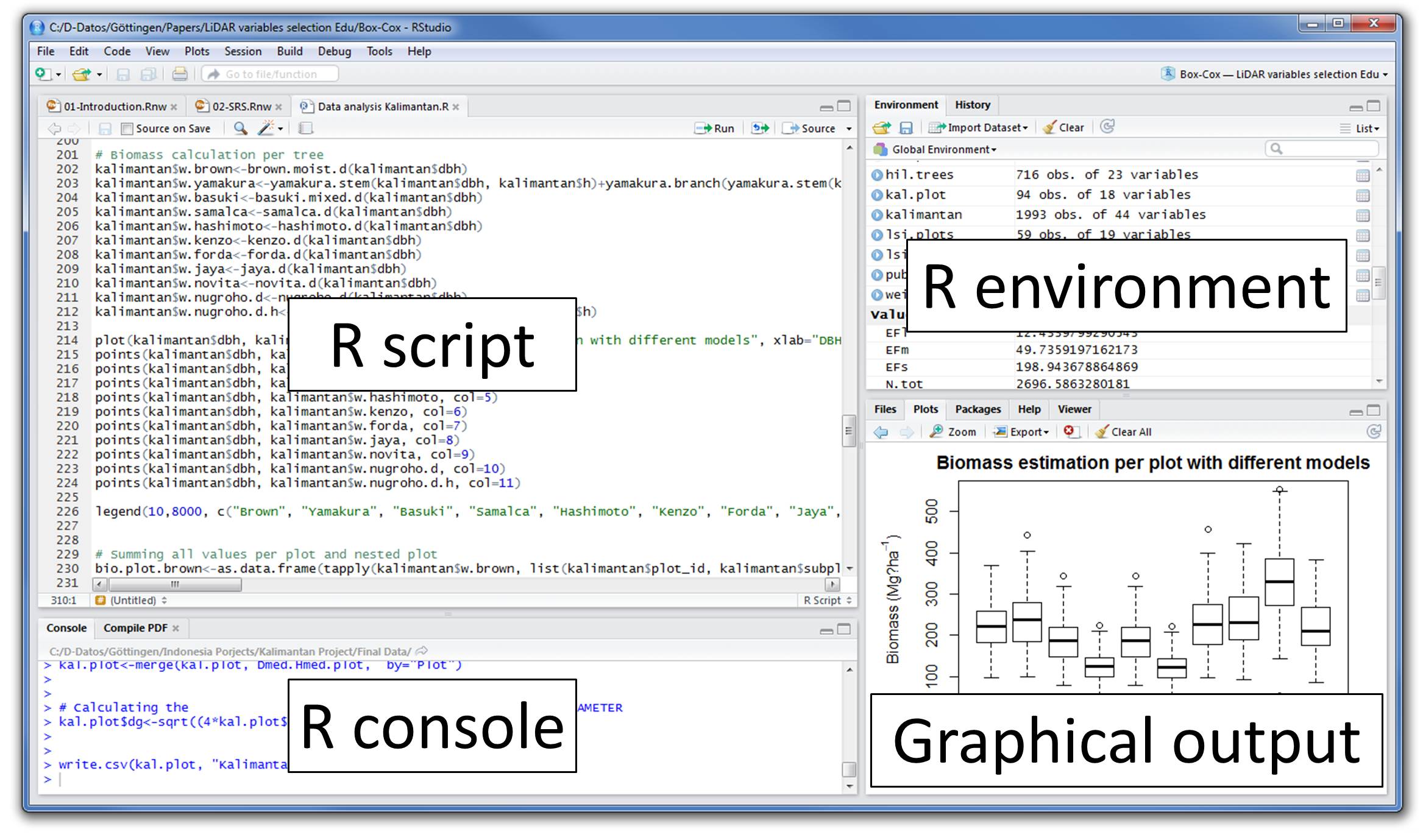
Let’s quickly understand the interface of R Studio:
- R Console: This area shows the output of code you run. Also, you can directly write codes in console. Code entered directly in R console cannot be traced later. This is where R script comes to use.
- R Script: As the name suggest, here you get space to write codes. To run those codes, simply select the line(s) of code and press Ctrl + Enter. Alternatively, you can click on little ‘Run’ button location at top right corner of R Script.
- R environment: This space displays the set of external elements added. This includes data set, variables, vectors, functions etc. To check if data has been loaded properly in R, always look at this area.
- Graphical Output: This space display the graphs created during exploratory data analysis. Not just graphs, you could select packages, seek help with embedded R’s official documentation.
How to install R Packages ?
The sheer power of R lies in its incredible packages. In R, most data handling tasks can be performed in 2 ways: Using R packages and R base functions. In this tutorial, I’ll also introduce you with the most handy and powerful R packages. To install a package, simply type:
install.packages("package name")
As a first time user, a pop might appear to select your CRAN mirror (country server), choose accordingly and press OK.
Note: You can type this either in console directly and press ‘Enter’ or in R script and click ‘Run’.
Basic Computations in R
Let’s begin with basics. To get familiar with R coding environment, start with some basic calculations. R console can be used as an interactive calculator too. Type the following in your console:
> 2 + 3
> 5
> 6 / 3
> 2
> (3*8)/(2*3)
> 4
> log(12)
> 1.07
> sqrt (121)
> 11
Similarly, you can experiment various combinations of calculations and get the results. In case, you want to obtain the previous calculation, this can be done in two ways. First, click in R console, and press ‘Up / Down Arrow’ key on your keyboard. This will activate the previously executed commands. Press Enter.
But, what if you have done too many calculations ? It would be too painful to scroll through every command and find it out. In such situations, creating variable is a helpful way.
In R, you can create a variable using <- or = sign. Let’s say I want to create a variable x to compute the sum of 7 and 8. I’ll write it as:
> x <- 8 + 7
> x
> 15
Once we create a variable, you no longer get the output directly (like calculator), unless you call the variable in the next line. Remember, variables can be alphabets, alphanumeric but not numeric. You can’t create numeric variables.
2. Essentials of R Programming
Understand and practice this section thoroughly. This is the building block of your R programming knowledge. If you get this right, you would face less trouble in debugging.
R has five basic or ‘atomic’ classes of objects. Wait, what is an object ?
Everything you see or create in R is an object. A vector, matrix, data frame, even a variable is an object. R treats it that way. So, R has 5 basic classes of objects. This includes:
- Character
- Numeric (Real Numbers)
- Integer (Whole Numbers)
- Complex
- Logical (True / False)
Since these classes are self-explanatory by names, I wouldn’t elaborate on that. These classes have attributes. Think of attributes as their ‘identifier’, a name or number which aptly identifies them. An object can have following attributes:
- names, dimension names
- dimensions
- class
- length
Attributes of an object can be accessed using attributes() function. More on this coming in following section.
Let’s understand the concept of object and attributes practically. The most basic object in R is known as vector. You can create an empty vector using vector(). Remember, a vector contains object of same class.
For example: Let’s create vectors of different classes. We can create vector using c() or concatenate command also.
> a <- c(1.8, 4.5) #numeric
> b <- c(1 + 2i, 3 - 6i) #complex
> d <- c(23, 44) #integer
> e <- vector("logical", length = 5)
Similarly, you can create vector of various classes.
Data Types in R
R has various type of ‘data types’ which includes vector (numeric, integer etc), matrices, data frames and list. Let’s understand them one by one.
Vector: As mentioned above, a vector contains object of same class. But, you can mix objects of different classes too. When objects of different classes are mixed in a list, coercion occurs. This effect causes the objects of different types to ‘convert’ into one class. For example:
> qt <- c("Time", 24, "October", TRUE, 3.33) #character
> ab <- c(TRUE, 24) #numeric
> cd <- c(2.5, "May") #character
To check the class of any object, use class(“vector name”) function.
> class(qt)
"character"
To convert the class of a vector, you can use as. command.
> bar <- 0:5
> class(bar)
> "integer"
> as.numeric(bar)
> class(bar)
> "numeric"
> as.character(bar)
> class(bar)
> "character"
Similarly, you can change the class of any vector. But, you should pay attention here. If you try to convert a “character” vector to “numeric” , NAs will be introduced. Hence, you should be careful to use this command.
List: A list is a special type of vector which contain elements of different data types. For example:
> my_list <- list(22, "ab", TRUE, 1 + 2i)
> my_list
[[1]]
[1] 22
[[2]]
[1] "ab"
[[3]]
[1] TRUE
[[4]]
[1] 1+2i
As you can see, the output of a list is different from a vector. This is because, all the objects are of different types. The double bracket [[1]] shows the index of first element and so on. Hence, you can easily extract the element of lists depending on their index. Like this:
> my_list[[3]]
> [1] TRUE
You can use [] single bracket too. But, that would return the list element with its index number, instead of the result above. Like this:
> my_list[3]
> [[1]]
[1] TRUE
Matrices: When a vector is introduced with row and column i.e. a dimension attribute, it becomes a matrix. A matrix is represented by set of rows and columns. It is a 2 dimensional data structure. It consist of elements of same class. Let’s create a matrix of 3 rows and 2 columns:
> my_matrix <- matrix(1:6, nrow=3, ncol=2)
> my_matrix
[,1] [,2]
[1,] 1 4
[2,] 2 5
[3,] 3 6
> dim(my_matrix)
[1] 3 2
> attributes(my_matrix)
$dim
[1] 3 2
As you can see, the dimensions of a matrix can be obtained using either dim() or attributes() command. To extract a particular element from a matrix, simply use the index shown above. For example(try this at your end):
> my_matrix[,2] #extracts second column
> my_matrix[,1] #extracts first column
> my_matrix[2,] #extracts second row
> my_matrix[1,] #extracts first row
As an interesting fact, you can also create a matrix from a vector. All you need to do is, assign dimension dim() later. Like this:
> age <- c(23, 44, 15, 12, 31, 16)
> age
[1] 23 44 15 12 31 16
> dim(age) <- c(2,3)
> age
[,1] [,2] [,3]
[1,] 23 15 31
[2,] 44 12 16
> class(age)
[1] "matrix"
You can also join two vectors using cbind() and rbind() functions. But, make sure that both vectors have same number of elements. If not, it will return NA values.
> x <- c(1, 2, 3, 4, 5, 6)
> y <- c(20, 30, 40, 50, 60)
> cbind(x, y)
> cbind(x, y)
x y
[1,] 1 20
[2,] 2 30
[3,] 3 40
[4,] 4 50
[5,] 5 60
[6,] 6 70
> class(cbind(x, y))
[1] “matrix”
Data Frame: This is the most commonly used member of data types family. It is used to store tabular data. It is different from matrix. In a matrix, every element must have same class. But, in a data frame, you can put list of vectors containing different classes. This means, every column of a data frame acts like a list. Every time you will read data in R, it will be stored in the form of a data frame. Hence, it is important to understand the majorly used commands on data frame:
> df <- data.frame(name = c("ash","jane","paul","mark"), score = c(67,56,87,91))
> df
name score
1 ash 67
2 jane 56
3 paul 87
4 mark 91
> dim(df)
[1] 4 2
> str(df)
'data.frame': 4 obs. of 2 variables:
$ name : Factor w/ 4 levels "ash","jane","mark",..: 1 2 4 3
$ score: num 67 56 87 91
> nrow(df)
[1] 4
> ncol(df)
[1] 2
Let’s understand the code above. df is the name of data frame. dim() returns the dimension of data frame as 4 rows and 2 columns. str() returns the structure of a data frame i.e. the list of variables stored in the data frame. nrow() and ncol() return the number of rows and number of columns in a data set respectively.
Here you see “name” is a factor variable and “score” is numeric. In data science, a variable can be categorized into two types: Continuous and Categorical.
Continuous variables are those which can take any form such as 1, 2, 3.5, 4.66 etc. Categorical variables are those which takes only discrete values such as 2, 5, 11, 15 etc. In R, categorical values are represented by factors. In df, name is a factor variable having 4 unique levels. Factor or categorical variable are specially treated in a data set. For more explanation, click here. Similarly, you can find techniques to deal with continuous variables here.
Let’s now understand the concept of missing values in R. This is one of the most painful yet crucial part of predictive modeling. You must be aware of all techniques to deal with them. The complete explanation on such techniques is provided here.
Missing values in R are represented by NA and NaN. Now we’ll check if a data set has missing values (using the same data frame df).
> df[1:2,2] <- NA #injecting NA at 1st, 2nd row and 2nd column of df
> df
name score
1 ash NA
2 jane NA
3 paul 87
4 mark 91
> is.na(df) #checks the entire data set for NAs and return logical output
name score
[1,] FALSE TRUE
[2,] FALSE TRUE
[3,] FALSE FALSE
[4,] FALSE FALSE
> table(is.na(df)) #returns a table of logical output
FALSE TRUE
6 2
> df[!complete.cases(df),] #returns the list of rows having missing values
name score
1 ash NA
2 jane NA
Missing values hinder normal calculations in a data set. For example, let’s say, we want to compute the mean of score. Since there are two missing values, it can’t be done directly. Let’s see:
mean(df$score)
[1] NA
> mean(df$score, na.rm = TRUE)
[1] 89
The use of na.rm = TRUE parameter tells R to ignore the NAs and compute the mean of remaining values in the selected column (score). To remove rows with NA values in a data frame, you can use na.omit:
> new_df <- na.omit(df)
> new_df
name score
3 paul 87
4 mark 91
Control Structures in R
As the name suggest, a control structure ‘controls’ the flow of code / commands written inside a function. A function is a set of multiple commands written to automate a repetitive coding task.
For example: You have 10 data sets. You want to find the mean of ‘Age’ column present in every data set. This can be done in 2 ways: either you write the code to compute mean 10 times or you simply create a function and pass the data set to it.
Let’s understand the control structures in R with simple examples:
if, else – This structure is used to test a condition. Below is the syntax:
if (<condition>){
##do something
} else {
##do something
}
Example
#initialize a variable
N <- 10
#check if this variable * 5 is > 40
if (N * 5 > 40){
print("This is easy!")
} else {
print ("It's not easy!")
}
[1] "This is easy!"
for – This structure is used when a loop is to be executed fixed number of times. It is commonly used for iterating over the elements of an object (list, vector). Below is the syntax:
for (<search condition>){
#do something
}
Example
#initialize a vector
y <- c(99,45,34,65,76,23)
#print the first 4 numbers of this vector
for(i in 1:4){
print (y[i])
}
[1] 99
[1] 45
[1] 34
[1] 65
while – It begins by testing a condition, and executes only if the condition is found to be true. Once the loop is executed, the condition is tested again. Hence, it’s necessary to alter the condition such that the loop doesn’t go infinity. Below is the syntax:
#initialize a condition
Age <- 12
#check if age is less than 17
while(Age < 17){
print(Age)
Age <- Age + 1 #Once the loop is executed, this code breaks the loop
}
[1] 12
[1] 13
[1] 14
[1] 15
[1] 16
There are other control structures as well but are less frequently used than explained above. Those structures are:
- repeat – It executes an infinite loop
- break – It breaks the execution of a loop
- next – It allows to skip an iteration in a loop
- return – It help to exit a function
Note: If you find the section ‘control structures’ difficult to understand, not to worry. R is supported by various packages to compliment the work done by control structures.
Useful R Packages
Out of ~7800 packages listed on CRAN, I’ve listed some of the most powerful and commonly used packages in predictive modeling in this article. Since, I’ve already explained the method of installing packages, you can go ahead and install them now. Sooner or later you’ll need them.
Importing Data: R offers wide range of packages for importing data available in any format such as .txt, .csv, .json, .sql etc. To import large files of data quickly, it is advisable to install and use data.table, readr, RMySQL, sqldf, jsonlite.
Data Visualization: R has in built plotting commands as well. They are good to create simple graphs. But, becomes complex when it comes to creating advanced graphics. Hence, you should install ggplot2.
Data Manipulation: R has a fantastic collection of packages for data manipulation. These packages allows you to do basic & advanced computations quickly. These packages are dplyr, plyr, tidyr, lubridate, stringr. Check out this complete tutorial on data manipulation packages in R.
Modeling / Machine Learning: For modeling, caret package in R is powerful enough to cater to every need for creating machine learning model. However, you can install packages algorithms wise such as randomForest, rpart, gbm etc
Note: I’ve only mentioned the commonly used packages. You might like to check this interesting infographic on complete list of useful R packages.
Till here, you became familiar with the basic work style in R and its associated components. From next section, we’ll begin with predictive modeling. But before you proceed. I want you to practice, what you’ve learnt till here.
Practice Assignment: As a part of this assignment, install ‘swirl’ package in package. Then type, library(swirl) to initiate the package. And, complete this interactive R tutorial. If you have followed this article thoroughly, this assignment should be an easy task for you!
3. Exploratory Data Analysis in R
From this section onwards, we’ll dive deep into various stages of predictive modeling. Hence, make sure you understand every aspect of this section. In case you find anything difficult to understand, ask me in the comments section below.
Data Exploration is a crucial stage of predictive model. You can’t build great and practical models unless you learn to explore the data from begin to end. This stage forms a concrete foundation for data manipulation (the very next stage). Let’s understand it in R.
In this tutorial, I’ve taken the data set from Big Mart Sales Prediction. Before we start, you must get familiar with these terms:
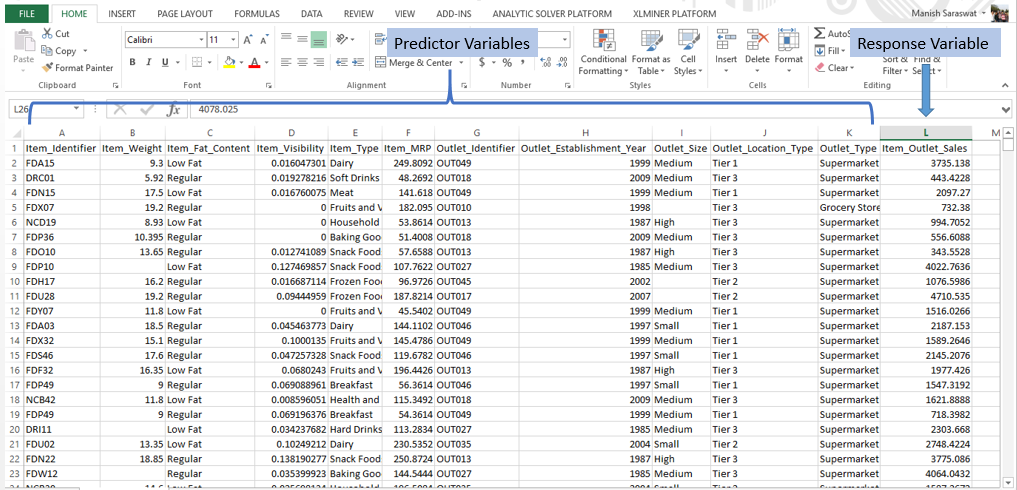
Response Variable (a.k.a Dependent Variable): In a data set, the response variable (y) is one on which we make predictions. In this case, we’ll predict ‘Item_Outlet_Sales’. (Refer to image shown below)
Predictor Variable (a.k.a Independent Variable): In a data set, predictor variables (Xi) are those using which the prediction is made on response variable. (Image below).
Train Data: The predictive model is always built on train data set. An intuitive way to identify the train data is, that it always has the ‘response variable’ included.
Test Data: Once the model is built, it’s accuracy is ‘tested’ on test data. This data always contains less number of observations than train data set. Also, it does not include ‘response variable’.
Right now, you should download the data set. Take a good look at train and test data. Cross check the information shared above and then proceed.
Let’s now begin with importing and exploring data.
#working directory
path <- ".../Data/BigMartSales"
#set working directory
setwd(path)
As a beginner, I’ll advise you to keep the train and test files in your working directly to avoid unnecessary directory troubles. Once the directory is set, we can easily import the .csv files using commands below.
#Load Datasets
train <- read.csv("Train_UWu5bXk.csv")
test <- read.csv("Test_u94Q5KV.csv")
In fact, even prior to loading data in R, it’s a good practice to look at the data in Excel. This helps in strategizing the complete prediction modeling process. To check if the data set has been loaded successfully, look at R environment. The data can be seen there. Let’s explore the data quickly.
#check dimesions ( number of row & columns) in data set
> dim(train)
[1] 8523 12
> dim(test)
[1] 5681 11
We have 8523 rows and 12 columns in train data set and 5681 rows and 11 columns in data set. This makes sense. Test data should always have one column less (mentioned above right?). Let’s get deeper in train data set now.
#check the variables and their types in train
> str(train)
'data.frame': 8523 obs. of 12 variables:
$ Item_Identifier : Factor w/ 1559 levels "DRA12","DRA24",..: 157 9 663 1122 1298 759 697 739 441 991 ...
$ Item_Weight : num 9.3 5.92 17.5 19.2 8.93 ...
$ Item_Fat_Content : Factor w/ 5 levels "LF","low fat",..: 3 5 3 5 3 5 5 3 5 5 ...
$ Item_Visibility : num 0.016 0.0193 0.0168 0 0 ...
$ Item_Type : Factor w/ 16 levels "Baking Goods",..: 5 15 11 7 10 1 14 14 6 6 ...
$ Item_MRP : num 249.8 48.3 141.6 182.1 53.9 ...
$ Outlet_Identifier : Factor w/ 10 levels "OUT010","OUT013",..: 10 4 10 1 2 4 2 6 8 3 ...
$ Outlet_Establishment_Year: int 1999 2009 1999 1998 1987 2009 1987 1985 2002 2007 ...
$ Outlet_Size : Factor w/ 4 levels "","High","Medium",..: 3 3 3 1 2 3 2 3 1 1 ...
$ Outlet_Location_Type : Factor w/ 3 levels "Tier 1","Tier 2",..: 1 3 1 3 3 3 3 3 2 2 ...
$ Outlet_Type : Factor w/ 4 levels "Grocery Store",..: 2 3 2 1 2 3 2 4 2 2 ...
$ Item_Outlet_Sales : num 3735 443 2097 732 995 ...
Let’s do some quick data exploration.
To begin with, I’ll first check if this data has missing values. This can be done by using:
> table(is.na(train))
FALSE TRUE
100813 1463
In train data set, we have 1463 missing values. Let’s check the variables in which these values are missing. It’s important to find and locate these missing values. Many data scientists have repeatedly advised beginners to pay close attention to missing value in data exploration stages.
> colSums(is.na(train))
Item_Identifier Item_Weight
0 1463
Item_Fat_Content Item_Visibility
0 0
Item_Type Item_MRP
0 0
Outlet_Identifier Outlet_Establishment_Year
0 0
Outlet_Size Outlet_Location_Type
0 0
Outlet_Type Item_Outlet_Sales
0 0
Hence, we see that column Item_Weight has 1463 missing values. Let’s get more inferences from this data.
> summary(train)
Here are some quick inferences drawn from variables in train data set:
- Item_Fat_Content has mis-matched factor levels.
- Minimum value of item_visibility is 0. Practically, this is not possible. If an item occupies shelf space in a grocery store, it ought to have some visibility. We’ll treat all 0’s as missing values.
- Item_Weight has 1463 missing values (already explained above).
- Outlet_Size has a unmatched factor levels.
These inference will help us in treating these variable more accurately.
Graphical Representation of Variables
I’m sure you would understand these variables better when explained visually. Using graphs, we can analyze the data in 2 ways: Univariate Analysis and Bivariate Analysis.
Univariate analysis is done with one variable. Bivariate analysis is done with two variables. Univariate analysis is a lot easy to do. Hence, I’ll skip that part here. I’d recommend you to try it at your end. Let’s now experiment doing bivariate analysis and carve out hidden insights.
For visualization, I’ll use ggplot2 package. These graphs would help us understand the distribution and frequency of variables in the data set.
> ggplot(train, aes(x= Item_Visibility, y = Item_Outlet_Sales)) + geom_point(size = 2.5, color="navy") + xlab("Item Visibility") + ylab("Item Outlet Sales") + ggtitle("Item Visibility vs Item Outlet Sales")

We can see that majority of sales has been obtained from products having visibility less than 0.2. This suggests that item_visibility < 2 must be an important factor in determining sales. Let’s plot few more interesting graphs and explore such hidden stories.
> ggplot(train, aes(Outlet_Identifier, Item_Outlet_Sales)) + geom_bar(stat = "identity", color = "purple") +theme(axis.text.x = element_text(angle = 70, vjust = 0.5, color = "black")) + ggtitle("Outlets vs Total Sales") + theme_bw()
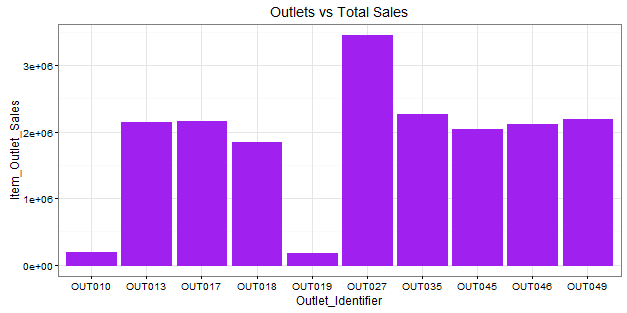
Here, we infer that OUT027 has contributed to majority of sales followed by OUT35. OUT10 and OUT19 have probably the least footfall, thereby contributing to the least outlet sales.
> ggplot(train, aes(Item_Type, Item_Outlet_Sales)) + geom_bar( stat = "identity") +theme(axis.text.x = element_text(angle = 70, vjust = 0.5, color = "navy")) + xlab("Item Type") + ylab("Item Outlet Sales")+ggtitle("Item Type vs Sales")
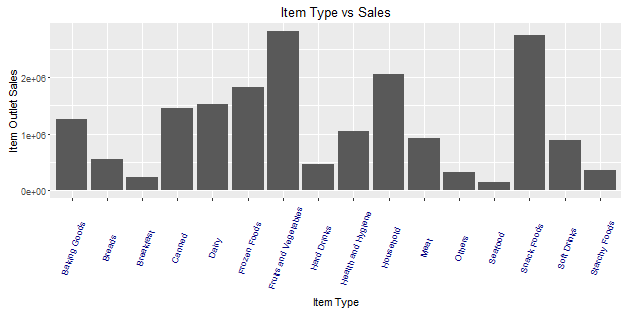
From this graph, we can infer that Fruits and Vegetables contribute to the highest amount of outlet sales followed by snack foods and household products. This information can also be represented using a box plot chart. The benefit of using a box plot is, you get to see the outlier and mean deviation of corresponding levels of a variable (shown below).
> ggplot(train, aes(Item_Type, Item_MRP)) +geom_boxplot() +ggtitle("Box Plot") + theme(axis.text.x = element_text(angle = 70, vjust = 0.5, color = "red")) + xlab("Item Type") + ylab("Item MRP") + ggtitle("Item Type vs Item MRP")
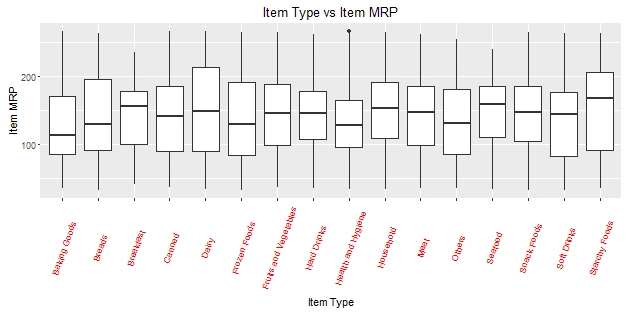

The black point you see, is an outlier. The mid line you see in the box, is the mean value of each item type. To know more about boxplots, check this tutorial.
Now, we have an idea of the variables and their importance on response variable. Let’s now move back to where we started. Missing values. Now we’ll impute the missing values.
We saw variable Item_Weight has missing values. Item_Weight is an continuous variable. Hence, in this case we can impute missing values with mean / median of item_weight. These are the most commonly used methods of imputing missing value. To explore other methods of this techniques, check out this tutorial.
Let’s first combine the data sets. This will save our time as we don’t need to write separate codes for train and test data sets. To combine the two data frames, we must make sure that they have equal columns, which is not the case.
> dim(train)
[1] 8523 12
> dim(test)
[1] 5681 11
Test data set has one less column (response variable). Let’s first add the column. We can give this column any value. An intuitive approach would be to extract the mean value of sales from train data set and use it as placeholder for test variable Item _Outlet_ Sales. Anyways, let’s make it simple for now. I’ve taken a value 1. Now, we’ll combine the data sets.
> test$Item_Outlet_Sales <- 1
> combi <- rbind(train, test)
Impute missing value by median. I’m using median because it is known to be highly robust to outliers. Moreover, for this problem, our evaluation metric is RMSE which is also highly affected by outliers. Hence, median is better in this case.
> combi$Item_Weight[is.na(combi$Item_Weight)] <- median(combi$Item_Weight, na.rm = TRUE)
> table(is.na(combi$Item_Weight))
FALSE
14204
Trouble with Continuous Variables & Categorical Variables
It’s important to learn to deal with continuous and categorical variables separately in a data set. In other words, they need special attention. In this data set, we have only 3 continuous variables and rest are categorical in nature. If you are still confused, I’ll suggest you to once again look at the data set using str() and proceed.
Let’s take up Item_Visibility. In the graph above, we saw item visibility has zero value also, which is practically not feasible. Hence, we’ll consider it as a missing value and once again make the imputation using median.
> combi$Item_Visibility <- ifelse(combi$Item_Visibility == 0,
median(combi$Item_Visibility), combi$Item_Visibility)
Let’s proceed to categorical variables now. During exploration, we saw there are mis-matched levels in variables which needs to be corrected.
> levels(combi$Outlet_Size)[1] <- "Other"
> library(plyr)
> combi$Item_Fat_Content <- revalue(combi$Item_Fat_Content,
c("LF" = "Low Fat", "reg" = "Regular"))
> combi$Item_Fat_Content <- revalue(combi$Item_Fat_Content, c("low fat" = "Low Fat"))
> table(combi$Item_Fat_Content)
Low Fat Regular
9185 5019
Using the commands above, I’ve assigned the name ‘Other’ to unnamed level in Outlet_Size variable. Rest, I’ve simply renamed the various levels of Item_Fat_Content.
4. Data Manipulation in R
Let’s call it as, the advanced level of data exploration. In this section we’ll practically learn about feature engineering and other useful aspects.
Feature Engineering: This component separates an intelligent data scientist from a technically enabled data scientist. You might have access to large machines to run heavy computations and algorithms, but the power delivered by new features, just can’t be matched. We create new variables to extract and provide as much ‘new’ information to the model, to help it make accurate predictions.
If you have been thinking all this time, great. But now is the time to think deeper. Look at the data set and ask yourself, what else (factor) could influence Item_Outlet_Sales ? Anyhow, the answer is below. But, I want you to try it out first, before scrolling down.
1. Count of Outlet Identifiers – There are 10 unique outlets in this data. This variable will give us information on count of outlets in the data set. More the number of counts of an outlet, chances are more will be the sales contributed by it.
> library(dplyr)
> a <- combi%>%
group_by(Outlet_Identifier)%>%
tally()
> head(a)
Source: local data frame [6 x 2]
Outlet_Identifier n
(fctr) (int)
1 OUT010 925
2 OUT013 1553
3 OUT017 1543
4 OUT018 1546
5 OUT019 880
6 OUT027 1559
> names(a)[2] <- "Outlet_Count"
> combi <- full_join(a, combi, by = "Outlet_Identifier")
As you can see, dplyr package makes data manipulation quite effortless. You no longer need to write long function. In the code above, I’ve simply stored the new data frame in a variable a. Later, the new column Outlet_Count is added in our original ‘combi’ data set. To know more about dplyr, follow this tutorial.
2. Count of Item Identifiers – Similarly, we can compute count of item identifiers too. It’s a good practice to fetch more information from unique ID variables using their count. This will help us to understand, which outlet has maximum frequency.
> b <- combi%>%
group_by(Item_Identifier)%>%
tally()
> names(b)[2] <- "Item_Count"
> head (b)
Item_Identifier Item_Count
(fctr) (int)
1 DRA12 9
2 DRA24 10
3 DRA59 10
4 DRB01 8
5 DRB13 9
6 DRB24 8
> combi <- merge(b, combi, by = “Item_Identifier”)
3. Outlet Years – This variable represent the information of existence of a particular outlet since year 2013. Why just 2013? You’ll find the answer in problem statement here. My hypothesis is, older the outlet, more footfall, large base of loyal customers and larger the outlet sales.
> c <- combi%>%
select(Outlet_Establishment_Year)%>%
mutate(Outlet_Year = 2013 - combi$Outlet_Establishment_Year)
> head(c)
Outlet_Establishment_Year Outlet_Year
1 1999 14
2 2009 4
3 1999 14
4 1998 15
5 1987 26
6 2009 4
> combi <- full_join(c, combi)
This suggests that outlets established in 1999 were 14 years old in 2013 and so on.
4. Item Type New – Now, pay attention to Item_Identifiers. We are about to discover a new trend. Look carefully, there is a pattern in the identifiers starting with “FD”,”DR”,”NC”. Now, check the corresponding Item_Types to these identifiers in the data set. You’ll discover, items corresponding to “DR”, are mostly eatables. Items corresponding to “FD”, are drinks. And, item corresponding to “NC”, are products which can’t be consumed, let’s call them non-consumable. Let’s extract these variables into a new variable representing their counts.
Here I’ll use substr(), gsub() function to extract and rename the variables respectively.
> q <- substr(combi$Item_Identifier,1,2)
> q <- gsub("FD","Food",q)
> q <- gsub("DR","Drinks",q)
> q <- gsub("NC","Non-Consumable",q)
> table(q)
Drinks Food Non-Consumable
1317 10201 2686
Let’s now add this information in our data set with a variable name ‘Item_Type_New.
> combi$Item_Type_New <- q
I’ll leave the rest of feature engineering intuition to you. You can think of more variables which could add more information to the model. But make sure, the variable aren’t correlated. Since, they are emanating from a same set of variable, there is a high chance for them to be correlated. You can check the same in R using cor() function.
Label Encoding and One Hot Encoding
Just, one last aspect of feature engineering left. Label Encoding and One Hot Encoding.
Label Encoding, in simple words, is the practice of numerically encoding (replacing) different levels of a categorical variables. For example: In our data set, the variable Item_Fat_Content has 2 levels: Low Fat and Regular. So, we’ll encode Low Fat as 0 and Regular as 1. This will help us convert a factor variable in numeric variable. This can be simply done using if else statement in R.
> combi$Item_Fat_Content <- ifelse(combi$Item_Fat_Content == "Regular",1,0)
One Hot Encoding, in simple words, is the splitting a categorical variable into its unique levels, and eventually removing the original variable from data set. Confused ? Here’s an example: Let’s take any categorical variable, say, Outlet_ Location_Type. It has 3 levels. One hot encoding of this variable, will create 3 different variables consisting of 1s and 0s. 1s will represent the existence of variable and 0s will represent non-existence of variable. Let look at a sample:
> sample <- select(combi, Outlet_Location_Type)
> demo_sample <- data.frame(model.matrix(~.-1,sample))
> head(demo_sample)
Outlet_Location_TypeTier.1 Outlet_Location_TypeTier.2 Outlet_Location_TypeTier.3
1 1 0 0
2 0 0 1
3 1 0 0
4 0 0 1
5 0 0 1
6 0 0 1
model.matrix creates a matrix of encoded variables. ~. -1 tells R, to encode all variables in the data frame, but suppress the intercept. So, what will happen if you don’t write -1 ? model.matrix will skip the first level of the factor, thereby resulting in just 2 out of 3 factor levels (loss of information).
This was the demonstration of one hot encoding. Hope you have understood the concept now. Let’s now apply this technique to all categorical variables in our data set (excluding ID variable).
>library(dummies)
>combi <- dummy.data.frame(combi, names = c('Outlet_Size','Outlet_Location_Type','Outlet_Type', 'Item_Type_New'), sep='_')
With this, I have shared 2 different methods of performing one hot encoding in R. Let’s check if encoding has been done.
> str (combi)
$ Outlet_Size_Other : int 0 1 1 0 1 0 0 0 0 0 ...
$ Outlet_Size_High : int 0 0 0 1 0 0 0 0 0 0 ...
$ Outlet_Size_Medium : int 1 0 0 0 0 0 1 1 0 1 ...
$ Outlet_Size_Small : int 0 0 0 0 0 1 0 0 1 0 ...
$ Outlet_Location_Type_Tier 1 : int 1 0 0 0 0 0 0 0 1 0 ...
$ Outlet_Location_Type_Tier 2 : int 0 1 0 0 1 1 0 0 0 0 ...
$ Outlet_Location_Type_Tier 3 : int 0 0 1 1 0 0 1 1 0 1 ...
$ Outlet_Type_Grocery Store : int 0 0 1 0 0 0 0 0 0 0 ...
$ Outlet_Type_Supermarket Type1: int 1 1 0 1 1 1 0 0 1 0 ...
$ Outlet_Type_Supermarket Type2: int 0 0 0 0 0 0 0 1 0 0 ...
$ Outlet_Type_Supermarket Type3: int 0 0 0 0 0 0 1 0 0 1 ...
$ Item_Outlet_Sales : num 1 3829 284 2553 2553 ...
$ Year : num 14 11 15 26 6 9 28 4 16 28 ...
$ Item_Type_New_Drinks : int 1 1 1 1 1 1 1 1 1 1 ...
$ Item_Type_New_Food : int 0 0 0 0 0 0 0 0 0 0 ...
$ Item_Type_New_Non-Consumable : int 0 0 0 0 0 0 0 0 0 0 ...
As you can see, after one hot encoding, the original variables are removed automatically from the data set.
5. Predictive Modeling using Machine Learning
Finally, we’ll drop the columns which have either been converted using other variables or are identifier variables. This can be accomplished using select from dplyr package.
> combi <- select(combi, -c(Item_Identifier, Outlet_Identifier, Item_Fat_Content, Outlet_Establishment_Year,Item_Type))
> str(combi)
In this section, I’ll cover Regression, Decision Trees and Random Forest. A detailed explanation of these algorithms is outside the scope of this article. These algorithms have been satisfactorily explained in our previous articles. I’ve provided the links for useful resources.
As you can see, we have encoded all our categorical variables. Now, this data set is good to take forward to modeling. Since, we started from Train and Test, let’s now divide the data sets.
> new_train <- combi[1:nrow(train),]
> new_test <- combi[-(1:nrow(train)),]
Linear (Multiple) Regression
Multiple Regression is used when response variable is continuous in nature and predictors are many. Had it been categorical, we would have used Logistic Regression. Before you proceed, sharpen your basics of Regression here.
Linear Regression takes following assumptions:
- There exists a linear relationship between response and predictor variables
- The predictor (independent) variables are not correlated with each other. Presence of collinearity leads to a phenomenon known as multicollinearity.
- The error terms are uncorrelated. Otherwise, it will lead to autocorrelation.
- Error terms must have constant variance. Non-constant variance leads to heteroskedasticity.
Let’s now build out first regression model on this data set. R uses lm() function for regression.
> linear_model <- lm(Item_Outlet_Sales ~ ., data = new_train)
> summary(linear_model)
Adjusted R² measures the goodness of fit of a regression model. Higher the R², better is the model. Our R² = 0.2085. It means we really did something drastically wrong. Let’s figure it out.
In our case, I could find our new variables aren’t helping much i.e. Item count, Outlet Count and Item_Type_New. Neither of these variables are significant. Significant variables are denoted by ‘*’ sign.
As we know, correlated predictor variables brings down the model accuracy. Let’s find out the amount of correlation present in our predictor variables. This can be simply calculated using:
> cor(new_train)
Alternatively, you can also use corrplot package for some fancy correlation plots. Scrolling through the long list of correlation coefficients, I could find a deadly correlation coefficient:
cor(new_train$Outlet_Count, new_train$`Outlet_Type_Grocery Store`)
[1] -0.9991203
Outlet_Count is highly correlated (negatively) with Outlet Type Grocery Store. Here are some problems I could find in this model:
- We have correlated predictor variables.
- We did one hot encoding and label encoding. That’s not necessary since linear regression handle categorical variables by creating dummy variables intrinsically.
- The new variables (item count, outlet count, item type new) created in feature engineering are not significant.
Let’s try to create a more robust regression model. This time, I’ll be using a building a simple model without encoding and new features. Below is the entire code:
#load directory
> path <- "C:/Users/manish/desktop/Data/February 2016"
> setwd(path)
#load data
> train <- read.csv("train_Big.csv")
> test <- read.csv("test_Big.csv")
#create a new variable in test file
> test$Item_Outlet_Sales <- 1
#combine train and test data
> combi <- rbind(train, test)
#impute missing value in Item_Weight
> combi$Item_Weight[is.na(combi$Item_Weight)] <- median(combi$Item_Weight, na.rm = TRUE)
#impute 0 in item_visibility
> combi$Item_Visibility <- ifelse(combi$Item_Visibility == 0, median(combi$Item_Visibility), combi$Item_Visibility)
#rename level in Outlet_Size
> levels(combi$Outlet_Size)[1] <- "Other"
#rename levels of Item_Fat_Content
> library(plyr)
> combi$Item_Fat_Content <- revalue(combi$Item_Fat_Content,c("LF" = "Low Fat", "reg" = "Regular"))
> combi$Item_Fat_Content <- revalue(combi$Item_Fat_Content, c("low fat" = "Low Fat"))
#create a new column 2013 - Year
> combi$Year <- 2013 - combi$Outlet_Establishment_Year
#drop variables not required in modeling
> library(dplyr)
> combi <- select(combi, -c(Item_Identifier, Outlet_Identifier, Outlet_Establishment_Year))
#divide data set
> new_train <- combi[1:nrow(train),]
> new_test <- combi[-(1:nrow(train)),]
#linear regression
> linear_model <- lm(Item_Outlet_Sales ~ ., data = new_train)
> summary(linear_model)
Now we have got R² = 0.5623. This teaches us that, sometimes all you need is simple thought process to get high accuracy. Quite a good improvement from previous model. Next, time when you work on any model, always remember to start with a simple model.
Let’s check out regression plot to find out more ways to improve this model.
> par(mfrow=c(2,2))
> plot(linear_model)
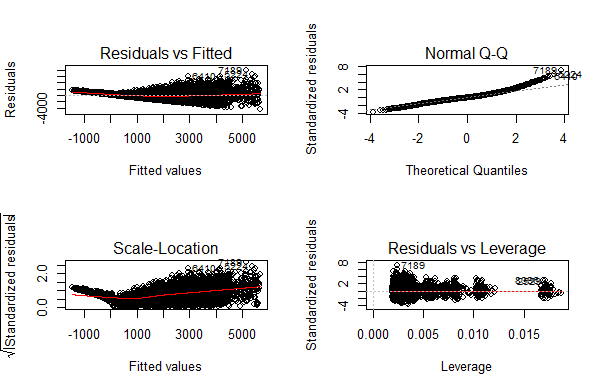
You can zoom these graphs in R Studio at your end. All these plots have a different story to tell. But the most important story is being portrayed by Residuals vs Fitted graph.
Residual values are the difference between actual and predicted outcome values. Fitted values are the predicted values. If you see carefully, you’ll discover it as a funnel shape graph (from right to left ). The shape of this graph suggests that our model is suffering from heteroskedasticity (unequal variance in error terms). Had there been constant variance, there would be no pattern visible in this graph.
A common practice to tackle heteroskedasticity is by taking the log of response variable. Let’s do it and check if we can get further improvement.
> linear_model <- lm(log(Item_Outlet_Sales) ~ ., data = new_train)
> summary(linear_model)
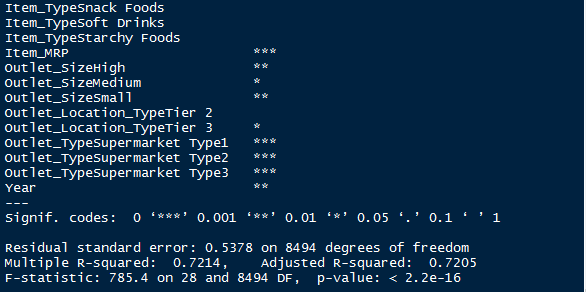
And, here’s a snapshot of my model output. Congrats! We have got an improved model with R² = 0.72. Now, we are on the right path. Once again you can check the residual plots (you might zoom it). You’ll find there is no longer a trend in residual vs fitted value plot.
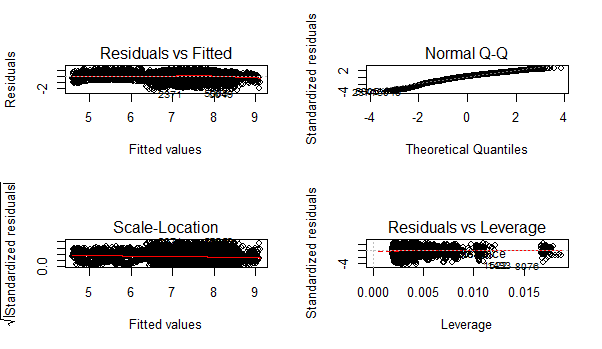
This model can be further improved by detecting outliers and high leverage points. For now, I leave that part to you! I shall write a separate post on mysteries of regression soon. For now, let’s check our RMSE so that we can compare it with other algorithms demonstrated below.
To calculate RMSE, we can load a package named Metrics.
> install.packages("Metrics")
> library(Metrics)
> rmse(new_train$Item_Outlet_Sales, exp(linear_model$fitted.values))
[1] 1140.004
Let’s proceed to decision tree algorithm and try to improve our RMSE score.
Decision Trees
Before you start, I’d recommend you to glance through the basics of decision tree algorithms. To understand what makes it superior than linear regression, check this tutorial Part 1 and Part 2.
In R, decision tree algorithm can be implemented using rpart package. In addition, we’ll use caret package for doing cross validation. Cross validation is a technique to build robust models which are not prone to overfitting. Read more about Cross Validation.
In R, decision tree uses a complexity parameter (cp). It measures the tradeoff between model complexity and accuracy on training set. A smaller cp will lead to a bigger tree, which might overfit the model. Conversely, a large cp value might underfit the model. Underfitting occurs when the model does not capture underlying trends properly. Let’s find out the optimum cp value for our model with 5 fold cross validation.
#loading required libraries
> library(rpart)
> library(e1071)
> library(rpart.plot)
> library(caret)
#setting the tree control parameters
> fitControl <- trainControl(method = "cv", number = 5)
> cartGrid <- expand.grid(.cp=(1:50)*0.01)
#decision tree
> tree_model <- train(Item_Outlet_Sales ~ ., data = new_train, method = "rpart", trControl = fitControl, tuneGrid = cartGrid)
> print(tree_model)
The final value for cp = 0.01. You can also check the table populated in console for more information. The model with cp = 0.01 has the least RMSE. Let’s now build a decision tree with 0.01 as complexity parameter.
> main_tree <- rpart(Item_Outlet_Sales ~ ., data = new_train, control = rpart.control(cp=0.01))
> prp(main_tree)
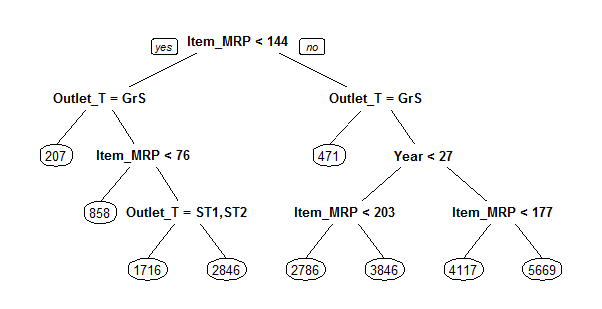
Here is the tree structure of our model. If you have gone through the basics, you would now understand that this algorithm has marked Item_MRP as the most important variable (being the root node). Let’s check the RMSE of this model and see if this is any better than regression.
> pre_score <- predict(main_tree, type = "vector")
> rmse(new_train$Item_Outlet_Sales, pre_score)
[1] 1102.774
As you can see, our RMSE has further improved from 1140 to 1102.77 with decision tree. To improve this score further, you can further tune the parameters for greater accuracy.
Random Forest
Random Forest is a powerful algorithm which holistically takes care of missing values, outliers and other non-linearities in the data set. It’s simply a collection of classification trees, hence the name ‘forest’. I’d suggest you to quickly refresh your basics of random forest with this tutorial.
In R, random forest algorithm can be implement using randomForest package. Again, we’ll use train package for cross validation and finding optimum value of model parameters.
For this problem, I’ll focus on two parameters of random forest. mtry and ntree. ntree is the number of trees to be grown in the forest. mtry is the number of variables taken at each node to build a tree. And, we’ll do a 5 fold cross validation.
Let’s do it!
#load randomForest library
> library(randomForest)
#set tuning parameters
> control <- trainControl(method = "cv", number = 5)
#random forest model
> rf_model <- train(Item_Outlet_Sales ~ ., data = new_train, method = "parRF", trControl = control, prox = TRUE, allowParallel = TRUE)
#check optimal parameters
> print(rf_model)
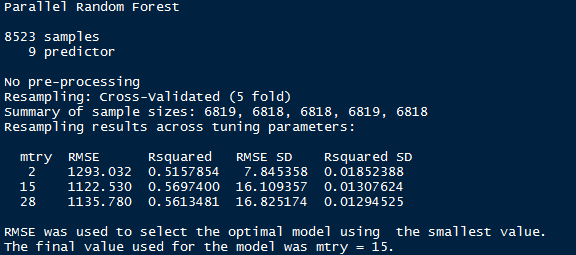
If you notice, you’ll see I’ve used method = “parRF”. This is parallel random forest. This is parallel implementation of random forest. This package causes your local machine to take less time in random forest computation. Alternatively, you can also use method = “rf” as a standard random forest function.
Now we’ve got the optimal value of mtry = 15. Let’s use 1000 trees for computation.
#random forest model
> forest_model <- randomForest(Item_Outlet_Sales ~ ., data = new_train, mtry = 15, ntree = 1000)
> print(forest_model)
> varImpPlot(forest_model)
This model throws RMSE = 1132.04 which is not an improvement over decision tree model. Random forest has a feature of presenting the important variables. We see that the most important variable is Item_MRP (also shown by decision tree algorithm).
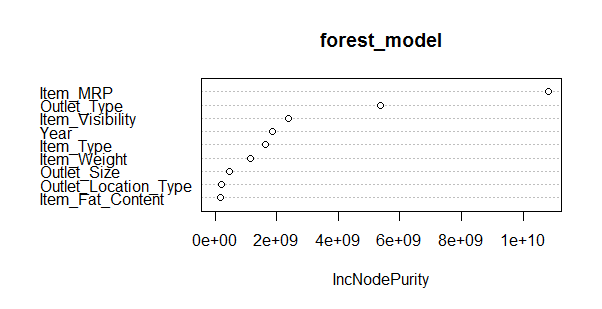
This model can be further improved by tuning parameters. Also, Let’s make out first submission with our best RMSE score by decision tree.
> main_predict <- predict(main_tree, newdata = new_test, type = "vector")
> sub_file <- data.frame(Item_Identifier = test$Item_Identifier, Outlet_Identifier = test$Outlet_Identifier, Item_Outlet_Sales = main_predict)
> write.csv(sub_file, 'Decision_tree_sales.csv')
When predicted on out of sample data, our RMSE has come out to be 1174.33. Here are some things you can do to improve this model further:
- Since we did not use encoding, I encourage you to use one hot encoding and label encoding for random forest model.
- Parameters Tuning will help.
- Use Gradient Boosting.
- Build an ensemble of these models. Read more about Ensemble Modeling.
Do implement the ideas suggested above and share your improvement in the comments section below. Currently, Rank 1 on Leaderboard has obtained RMSE score of 1137.71. Beat it!
End Notes
This brings us to the end of this tutorial. Regret for not so happy ending. But, I’ve given you enough hints to work on. The decision to not use encoded variables in the model, turned out to be beneficial until decision trees.
The motive of this tutorial was to get your started with predictive modeling in R. We learnt few uncanny things such as ‘build simple models’. Don’t jump towards building a complex model. Simple models give you benchmark score and a threshold to work with.
In this tutorial, I have demonstrated the steps used in predictive modeling in R. I’ve covered data exploration, data visualization, data manipulation and building models using Regression, Decision Trees and Random Forest algorithms.
Did you find this tutorial useful ? Are you facing any trouble at any stage of this tutorial ? Feel free to mention your doubts in the comments section below. Do share if you get a better score.
Edit: On visitor’s request, the PDF version of the tutorial is available for download. You need to create a log in account to download the PDF. Also, you can bookmark this page for future reference. Download Here.










Thanks for sharing! Can this content be available in a Pdf format? Thanks,
Welcome Steve. I can make that available. I'll email it to you shortly.
plz mail pdf on [email protected]
Nice writeup useful, thnaks Samue
Welcome Samuel !
Thanks Manish. You wrote an amazing article for beginners. I was looking for an article like this which clears the basics of R without refering to any books and all. Even I request you to send me the doc or pdf of this so that i can get it print to make it handy to read.
Thanks Himanshu ! PDF is available for download. Link is added at the end of tutorial.
good one. pl mail me a pdf as well
Hi Manish Could you please share the pdf with me as well. I am a starter in R and this can help as a compact guide for myself when trying out different things. Thanks
Hello, when I type log(12) I get 2.484907 as a result. What seems to be the problem ?
@RadMou, It seems that there is a typo in the article. The fact is: 'log uses base e' ; log10 uses base 10' and 'log2 uses base 2'. You can see that these commands print different values: log(12) # log to the base e log10(12) # log to the base 10 log2(12) # log to the base 2 Hope this helps.
Thanks Manish . would be grateful if can be made available in PDF .
Hi Zamin PDF is available for download.
Hi Manish, This is very helpful for beginners like me. Looking forward for more. Is there any way I can get this in PDF format? It would be really helpful My email id is [email protected]. Thank you very much!.
Thanks Manish. This is a great help! I have a questions - I noticed that R automatically takes care of the factor variables (by converting them to n or n-1 dummy variables) while performing linear regression. Do you recommend that we do it explicitly?
Hi Anish In case of linear regression, decision trees, random forest, kNN, it is not necessary to convert categorical variables explicitly as these algorithms intrinsically breaks a categorical variables with n - 1 levels. However, if you are using boosting algorithms (GBM, XGboost) it is recommended to encode categorical variables prior to modeling. On a similar note, if you have followed this tutorial you'll find that I started with one hot encoding and got a terrible regression accuracy. Later, I used the categorical variables as it as, and accuracy improved.
good presentation. can you please provide it in pdf format.
Very helpful for beginners, thanks a lot!!!! keep it up.
Welcome !
Manish, Very valuable tutorial. TY. If it is not too much of a trouble. Can you please make a PDF version as a link on the tutorial, please. Thanks. Regards Raman
Hi Raman I've added the PDF link at the end of this tutorial.
Thanks for sharing this article. This is really help to us. When I ran these script on Rstudio I got two errors for ggplot after I tried " install.packages("ggplot2") AND "install.packages('ggplot2',dependencies = TRUE) "and I got the following error > ggplot(train, aes(x= Item_Visibility, y = Item_Outlet_Sales)) + geom_point(size = 2.5, color="navy") + xlab("Item Visibility") + ylab("Item Outlet Sales") Error: could not find function "ggplot" And also for merge data > combi <- merge(b, combi, by = "Outlet_Identifier") Error in fix.by(by.x, x) : 'by' must specify a uniquely valid column Can you help me why this happen. Once again 'Thank You So Much' because I learn new things about R. Thanks, Atul
Hi Atul After installing the ggplot2 package, you should call the package in the next step using library(ggplot2). Then run the ggplot code, it should work. merge function is used from package plyr. Have you installed it ? Let me know.
can u share any material of data science
Erratum : I'm not sure if the problem is from my computer, but : - When I execute head(b) I get : DRA12 9 RA24 10 And not OUT027 2215.876 OUT035 1463.705 So the command combi <- merge(b, combi, by = "Outlet_Identifier") should be combi <- merge(b, combi, by = "Item_Identifier") instead - Also in head(c) there is a problem with the years, all rows are for 1985.
"Hence, we see that column Item_Visibility has 1463 missing values. Let’s get more inferences from this data." it's the Item_Weight variable that has missing values Also in "Label Encoding and One Hot Encoding" : the variable Item_Visibility has 2 levels: Low Fat and Regular It's Item_Fat_Content not Item_Visibility
Hi Thanks for pointing out. Made the changes. In head(c), I wanted to show that using the "mutate" command, count value of years get automatically aligned to their particular year value. Hence, I sorted it. For example, the year 1985 would get 25 as count value at all the places in count column. Anyways, I've put a better picture of year count now. Hope this helps.
Hi Manish, I am unable to download the pdf as i get a blank page. Kindly check
Thanks. Works now after i relogin
Hii, When I use full_join for Outlet Years my rowcount increase to 23590924. I did not understand why full join is used and why rowcount is increasing.
Hi Ambuj full_join function returns all rows and all columns from the chosen data sets. And, if a value is not present it blatantly returns NA. In your case, you might not have specified the "by" parameter in full_join.
What I did was after c which has 14204 rows as flws : d % group_by(Outlet_Establishment_Year)%>% distinct() then combi <- merge(d, combi, by = "Outlet_Establishment_Year") combi will now be ready for label encoding...
Dear Ambuj, After generated c .. i created d using distinct d % group_by(Outlet_Establishment_Year)%>% distinct() Then merge d with combi as flws : combi <- merge(d, combi, by = "Outlet_Establishment_Year") Then ready for encoding. Thanks
Hi , Can you please send me the pdf file on [email protected] as i am unable to download the file from the link provided? Thanks in advance
Hi Gaurav, As mentioned, you need to create a one-time user account to download the pdf. You can find the link in the End Notes.
Could you please share the data (..../Data/BigMartSales) that you have used here so that we can play it with ?
It seems that your PDF file is missing in the correct link. May I request you to update it. Thanks in advance....
I got the PDF file, Thanks...
nice tutorial. I have 2 questions so far a) how to save my work - for e.g all the data manipulation steps i did are lost the next day and i have to start from the setwd(path) command again b) what is the difference between merge and full_join in the tutorial? when is each command more appropriate? c) The group by Item_identifier is not working properly. The sample output is wrong
Hi Buvana Answer a ) Do you directly write codes in console ? Use R Studio. You should use R script as they can be saved in .R format and helps you to retrieve codes at later time. For more information, check the first section of this tutorial. Answer b) full_join is used when we wish to combine two columns. It return NA when no matching value are found. merge is used when we wish to combine two columns based on a column type. In full_join, you don't need to specify "by" parameter. Answer c) Thank for pointing out. Sorted now.
Hi, In the Random Forest section, could you please explain why did you use ntree = 1000 after finding mtry = 15? Cheers,
Hi Guilherme If you carefully check random forest section, I've initially done cross validation using caret package. Cross validation provided the optimal value of mtry and ntree at which the RMSE is least (check output). I, then used those parameters in the final random forest model. Another method to choose mtry and ntree is hit and trial, which is certainly time consuming and inconsistent. You may try this experiment at your end, and let me know if you obtain lesser RMSE than what I've got.
Thank you very much for this wonderful and unique post. i came to this site to participate "date with your data" competition. i was puzzled looking at the datsets like train,test and sample & i dont have any idea what,and how to solve this. later on i came across this post (thank God i did) and really after going through your post i gained confidence & i got a clear picture on how to handle these competitions. once agian thanx from bottom of my heart.since i m completely new to this i have few doubts... 1) in "linear_model <- lm(Item_Outlet_Sales ~ ., data = new_train)" what does tilde(~) followed by dot (.) means? 2) what is the best RMSE score for any model? 3) so both train and test datsets are same,only thing is test data doesnt have response variable. But, if we do know the response variable value from train dataset, again why we we are calculating it for test data set? is it because we want to construct a model which predicts the future outcomes, but we want to test how good our model predicts value, so thats why we took sample from main dataset and cross check our predicted values with that of main dataset ? correct me if my understanding is wrong...
Hi Arfath Good to know that you have started learning. Answer 1: tilde(~) followed by dot (.) tells the model to select all the variables at once. Otherwise, it would be so much inconvenient to write name of all variables one by one. Imagine the time which would get wasted if you have got 200 variables to write. Therefore, use this short sign tilde(~) followed by dot (.) Answer 2: Ideally, every model strives for achieve RMSE as much as close to Zero. Because, Zero means your model has accurately predicted the outcome. But, that's not possible. Since, every model has got irreducible error which affects the accuracy. Hence, the best RMSE score is the least score you can get. Answer 3: You are absolutely. Train data set has response variable and a model is trained on that. This model gives you a fantastic RMSE score. But, it is worthless until it predicts with same accuracy on out of sample data. The ultimate aim for this model is to make future predictions. Right ? Hence, test data is used to check out of sample accuracy of the model. If the accuracy is not as good as you achieved on train data set, it suggests that overfitting has taken place. I would recommend you to read Introduction to Statistical Learning. Download link is available in my previous article: http://www.analyticsvidhya.com/blog/2016/02/free-read-books-statistics-mathematics-data-science/
I am a little late to the game. How do i download the BigMartSales data?
Hi Vijay Link is available in the tutorial.
Thanks for sharing. I just can not understand what the One Hot Encoding means and how to use it. Because I just new here. Thanks!
Hi Alfa One Hot Encoding is nothing but, splitting the levels of a categorical variable into new variable. The new variables will be encoded with 0s and 1s. 1s represent the presence of information. 0s represent the absence of information. For example: Suppose, we have a variable named as Hair Color. It has 3 levels namely Red Hair, Black Hair, Brown Hair. Doing one hot encoding of this variable, will result in 3 different variables namely Red Hair, Black Hair, Brown Hair. And, the original variable Hair Color will be removed from data set. If someone has Red Hair, Red Hair variable will be 1, Black Hair will be 0, Brown Hair will be 0. If someone has Black Hair, Red Hair variable will be 0, Black Hair will be 1, Brown Hair will be 0. If someone has Brown Hair, Red Hair variable will be 0, Black Hair will be 0, Brown Hair will be 1. This is One Hot Encoding.
Can someone please mail me the data sets we need for this article to [email protected]. I couldn't find at the mentioned location. It would be really helpful. Thanks
Hi Midhun The data set will be available for download from tomorrow onwards (13th March 2016) Regret the inconvenience caused.
Hi Manish, It's a great article & gives a good start for beginner like me. Can you please share the data. I can't download it from the link as the contest is not active. Thank You
Hi Manoj The data set will be available for download from tomorrow onwards. (13th March 2016)
Good Day...When I try to instal library(swirl) n R studio console ,,it states its not found in the version R.3..2.4.. I got errors which states"Warning in install.packages : package ‘library(swirl)’ is not available (for R version 3.2.4)" Can somebody explain to me this peculiarity and how can I sort it out... Thanks
Hi Roy First you should install swirl package and then call it using library function. Use the commands below. > install.packages("swirl") > library(swirl)
Hi Manish, The datasets are available now. Thank you so much.
I encounter problems to log in http://datahack.analyticsvidhya.com/signup... Can you help me ? I want to log in to then download the data set... Thanks in advance
Hello There were some technical updates going on at the server. Things are fine now. You may try again. Regret the inconvenience caused.
trying feature engineering of the outlet _establishment year ,but the code for merging is creating a lot of rows , i tried both merge as well full join .
hello sir i am a fresher electrical engineer and my maths and logical thinking is good can i become data scientist sir give me some advice thanks
I did try to see the link to try the " Big Market Prediction" but unable to open it as it requires membership. Now when I apply for the analytics Vidhya membership by signing up I got and Invalid Request twice ... May I know how I can get over this issue.. Why I can't sign up..so I can continue with my R self tutorial work..
Hi Thanks for an amazing article. Can you please email me the data used.
Hi Hulisani Please download the data set from here: http://datahack.analyticsvidhya.com/contest/practice-problem-big-mart-sales-iii
Hi, I am facing a problem in Random Forest execution. I am using R Studio (R version 3.2.4 Revised) When I am trying to run the code; > rf_model print(rf_model), it is returning error in this form : Error in { : task 1 failed - "cannot allocate vector of size 554.2 Mb" In addition: Warning messages: 1: executing %dopar% sequentially: no parallel backend registered 2: In eval(expr, envir, enclos) : model fit failed for Fold1: mtry=15 Error in { : task 1 failed - "cannot allocate vector of size 354.7 Mb" 3: In eval(expr, envir, enclos) : model fit failed for Fold2: mtry= 2 Error in { : task 1 failed - "cannot allocate vector of size 177.3 Mb" 4: In eval(expr, envir, enclos) : model fit failed for Fold2: mtry=28 Error in { : task 1 failed - "cannot allocate vector of size 177.3 Mb" 5: In eval(expr, envir, enclos) : model fit failed for Fold3: mtry=15 Error in { : task 1 failed - "cannot allocate vector of size 177.4 Mb" 6: In eval(expr, envir, enclos) : model fit failed for Fold4: mtry= 2 Error in { : task 1 failed - "cannot allocate vector of size 354.8 Mb" 7: In eval(expr, envir, enclos) : model fit failed for Fold4: mtry=28 Error in { : task 1 failed - "cannot allocate vector of size 354.8 Mb" 8: In eval(expr, envir, enclos) : model fit failed for Fold5: mtry=15 Error in { : task 1 failed - "cannot allocate vector of size 177.4 Mb" 9: In nominalTrainWorkflow(x = x, y = y, wts = weights, info = trainInfo, : There were missing values in resampled performance measures. 10: display list redraw incomplete Timing stopped at: 1.26 0.3 2.49 Can you please suggest me any way out of this issue?
The code I am trying to run is : rf_model <- train(Item_Outlet_Sales ~ ., data = new_train, method = "parRF", trControl = control, prox = TRUE, allowParallel = TRUE) print(rf_model)
Hi Priyanka Had I been at your place, I wouldn't have experimented with parallel random forest on this problem. Why make things complicated when it can be done in a simple way! Also, make sure that you drop the ID column before running any algorithm. Things should work fine then.
Hi Manish, After reading the whole article, I feel u have done a great job and have given more than enough data for a beginner. I'm thankful to u for sharing all your solutions, this would give us different thought for us to start with. Regards, Raju.
Glad it helped you. Thanks for your kind words Raju! :)
Good morning I can not find the data set. Any suggestion?
Hi Gregory Please download the data from here: http://datahack.analyticsvidhya.com/contest/practice-problem-big-mart-sales-iii
OK. I've registered and I think it'll be OK. Thanks
I know this is months after this great article was published, but i'm just now working through this and the BigMart Sales Prediction dataset isn't available. Is it available elsewhere?
Hi Toddim, The data set is very well available. I've already updated the links. You can download the data from here: http://datahack.analyticsvidhya.com/contest/practice-problem-big-mart-sales-iii
Hi Manish, First of all thanks for a great article. I encountered with a issue when I was running the code- combi <- full_join(c, combi, by="Outlet_Establishment_Year") it is giving me error- Error: std::bad_alloc what it is and how to correct this...
2. combi <- dummy.data.frame(combi, names = c('Outlet_Size','Outlet_Location_Type','Outlet_Type', 'Item_Type_New'), sep='_') Error in sort.list(y) : 'x' must be atomic for 'sort.list' Have you called 'sort' on a list?
Solution of this problem is present here: https://stackoverflow.com/questions/49718950/error-in-sort-listy-x-must-be-atomic-for-sort-list-have-you-called-sort
Hi Manish, Can you please let me know what do you mean by Item_Fat_Content has mismatched factor levels?
Even I want to know, what do the author meant by "Item_Fat_Content has mismatched factor levels",
Hi Manish. Thanks for this article. Very well written and will help all. I have one query: I could follow your post very well before'Graphical representation of Variables', after which I am unable to figure out how to write these codes and what do they mean & signify, how to know which command to use & when? I am a beginner in R . Can you please suggest what to do in order for me to fully understand all the steps from 'Graphical Representation'. This includes Data manipulation and Predictive modeling as well. Thanks a lot.
Very great article and thank you so much for sharing your knowledge! I am not sure if others have some questions with me, but I list my questions. Hope you have some time to take a look at it. Thank you again. 1. About the difference between label encoding and one hot encoding. For label encoding, your example is convert the 2 levels variables item_Fat_Content into 0 and 1. If I have a variable US state (50 levels = 50 States), is it means I just need simply trans the states to number 1-50? But it is still a one variables, just from category to numerical, am I right? 2. For one hot encoding, I need split into 50 variables (50 States) and marked them as 0s and 1s to indicate existence or non-existence, am I right? 3. So what is the advantage and disadvantage to convert the category variables into numeric variables? Why do we need to do this transformation? 4. In the article it said, ‘We did one hot encoding and label encoding. That’s not necessary since linear regression handle categorical variables by creating dummy variables intrinsically.’ How do we know which model we need to do the one hot encoding/ label encoding? 5. You mentation correlated variables. What level of correlation we need to remove the correlated variables? 0.5 or 0.6 or 0.7 ? And if two variables is correlated, how to decide which one we should remove? Is there any standard about it? 6. I am running logistic regression, when I remove one of the correlated variables (0.68), the R² dropped, is it means this level (0.68) correlation is acceptable? 7. The liner regression model with funnel share means heteroscedasticity. So how to evaluate the logistic regression with Residuals vs Fitted graph? 8. In the article, it is said ‘This model can be further improved by detecting outliers and high leverage points.’ what is the technical to deal with these points? Just simply remove the record or use the average to replace the value or other ways? 9. ‘optimum cp value for our model with 5 fold cross validation.’ In my mind, cross validation is used for evaluate the model stability which is the last step. However, at here, we use cross validation to optimum cp value, am I understand right? 10. Why are you using 5 fold cross validation instead of 4 fold or 6 fold or 10 fold? 11. When I running the model, it always have error told me the tree cannot split. Is there any requirement with the decision tree? Such as we cannot use category variables in decision tree? 12. How to do the Parameters Tuning for random forest? Could you points any arterials? Thank you !!!
In the below excerpts of the article: ""Data Frame: This is the most commonly used member of data types family. It is used to store tabular data. It is different from matrix. In a matrix, every element must have same class. But, in a data frame, you can put list of vectors containing different classes. This means, every column of a data frame acts like a list. Every time you will read data in R, "" it seems bit unconvincing that column of a dataframe acts like a list, instead column it should be row as per my understanding:
just think of a dataframe as an excel table. should help in understanding dataframes first column could be student_id (combination of alphabets and integers e.g. 05EL01) second column could be first name (e.g. Monish) third column could be math score (e.g. 90) 1. each column individually will contain data of one single type 2. the columns might be of different types as compared to each other
Hi I am beginner in Data Science using R. I was going through your well articulated article on Data Science using R. I was practicing your Big Mart Predication and got confused with one step , where it checks the missing values in train data exploration. As per R and this tutorial , there is only missing values (i assume blank is being considered as missing data) in "Item_Weight" but data is also missing in "Outlet_Size" in Train CSV.. But neither R or this tutorial is showing "Outlet_Size" as missing values observations. Can you please let me know how and why "Outlet_Size" is not considered as missing values in data exploration of train.
not very sure, but i would guess its because "Item_Weight" has numerical values while "Outlet_Size" has categorical values. team, is this so or is there some other reason? please, let me also know at [email protected]
Hi I would also like to know what all mathematical concepts like algebra , statics, are required to learn Data Science using R? Can anybody list down all mathematical concepts required for Data Science? Thanks Vaibhav Gupta
Hello, I had an error when launching RStudio. I downloaded it again and installed it again, but when I downloaded for the second time I found this phrase: "RStudio requires R 2.11.1 (or higher). If you don't already have R, you can download it here." (here is a link) So, before installing this, it looks like normal R has to be installed first. I write this in case someone had the same problem. Good job with the web, I really like it :)
As someone who came from a non-coding background, you should know that small details can become HUGE hindrances in the learning process of a beginner. On the Essentials part of the article, this code doesn't work: > bar class(bar) > "integer" > as.numeric(bar) > class(bar) > "numeric" > as.character(bar) > class(bar) > "character" You have to actually set it as 'bar <- as.numeric(bar)' on the 4th line. Please, keep those small things in mind. It is insanely difficult for someone like me to learn this content, if things are any less than perfect, it really becomes impossible (I just spent almost an hour to figure out why I couldn't change the class of the object, and in the end, had to ask for external help since I couldn't troubleshoot it myself). Otherwise, great article, keep the great work up! Cheers.
Thanks you made R programming simpler. Could you please email the PDF of the same.
Hi Team Analytics Vidhya, Can you please provide the updated link to download the dataset. Thanks in advance.
Hii, When I use full_join for Outlet Years my rowcount increase to 23590924. I did not understand why full join is used and why rowcount is increasing. can you help me with the exact code, that I have to put while inserting data in c?
Hii, When I use full_join for Outlet Years my row count increase to 23590924. I did not understand why full join is used and why row count is increasing. can you help me with the exact code, that I have to put while inserting data in c?
Sorry it is not working. It is asking to register. Once registered asking to go to provide mail to activate account. Once I went to my email and click on the activation link sent by analyticsvidya.com I am getting page saying your account not identified. I am not able to download. please help.
With R language being widely adopted in many data science applications it becomes very important for data science enthusiasts to have this skill. While there are many guides on R, this particular guide is very insightful and easy to understand.
hi, can anybody tell, 1. how do i run my LM model on the test data set 2. how to convert the result into csv format for uploading to solution checker any other info required to upload to the solution checker?
Hi team, I am not able to login to your webiste (during downloading the R Pdf file). so could you please rectify it ? Meanwhile please mail me the pdf @ [email protected] Thanks and Regards, Vishal Sharma.
Hi, This article is of great help to those who are learning R from scratch. First, thanks for creating this article, I have some doubts. What is the difference between Categorical and Continous variable ? What is the difference between Categorical and Factor variable? Can you please elaborate more on the basics of these classification of the variable? Awaiting response. Thanks, Anindo
Hi, Thanks for the wonderful tutorial. Somehow, I am not able to download the .pdf file. I tried logging in multiple times. Could you please mail me the pdf file @ [email protected]. Thanks again!
R is very useful to us/all researchers and analysts
Hello Everyone, I have a very small doubt, 1) I went through some case studies where they have used string as factors as false,and later created dummy variables for all the categorical variables by using functions or averages.. 2)while in some case studies they used package dummies,or caret to do the same 3) we can also use one hot encoding for the same What's the difference in all these 3 techniques,and when to use them..and if we have packages create dummies ,why to use other techniques... Pls help .. Thanks in advance
Hello admin,i am unable to find dataset in pdf format even though i have an account help me
Thank you for such useful article. Can you recommend any better article related to "R to build neural networks"? Appreciate it.
Thank you very much!!! Very well explained, esay to follow...great JOB!!
RESPECTED SIR, I HAVE GOOD KNOWLEDGE OF PROGRAMMING AND TESTING… BUT THERE ARE MAJOR GAP IN MY EDUCATIONAL AND PROFESSIONAL YEARS. SO, CONSIDERING ME AS A FRESHER “DATA SCIENCE” IS GOOD FOR MY FUTURE GROWTH ? PLEASE HELP ME OUT…
Hello, Completed the above tutorial in 14 days, its really helping to boost my confidence in my work place as a data scientist. As I am totally new to this domain (working as a fresher), what else i need to learn so that I can improve my analytical skills ? I am already learning R language. What else I need to learn so as to become an effective data scientist ? Please Guide me !
Hi Toofan, You can follow the Learning path, here is the link.
For anyone here saying this is a great article, I would need you to take a closer look, and run each code yourself. Some lines of codes contain typos. I had the same issues that other people had in the previous comments. So I had to search for other materials to correct the codes. The biggest problem is, as I read and tried towards the end, when I did the random forest, the R took forever to run, and never returned the result. the codes for cross validation of random forest model is: # set tuning parameters control <- trainControl(method = "cv", number = 5) # random forest model rf_model <- train(Item_Outlet_Sales ~ ., data = new_train, method = "parRF", trControl = control, prox = TRUE, allowParallel = TRUE) The codes for building model with randomForest package is: forest_model <- randomForest(Item_Outlet_Sales ~ ., data = new_train, mtry = 15, ntree = 1000) All above codes were originated from the article, And the article further explained "This package causes your local machine to take less time in random forest computation". In fact, there is absolutely no way these codes will get random forest perform faster. Would you check your codes to make sure they were correct, and can you explain how did you build your random forest model with less processing time? Please demonstrate it with codes, not giving me a broad and general response would be much appreciated. Thank you.
Looks like the hackathon has ended. I checked the website many times and couldn't find it. Doesn't make sense to use this tutorial when the dataset is not available
Hi Janani, You can download the dataset from this link.
Hi! i have a question. In mtcars data set in R i want to gather 8th, and 9th column i.e (vs and am). how can we do that?
You are the best human being I have ever come across on earth. Thank you so much